Probe CUST_41258_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41258_PI426222305 | JHI_St_60k_v1 | DMT400077168 | ACTTGGCAGAGAAATTCAAGCCGCGAGATGAAGATAAAGCACTTTCTGAGGTCATTTCAG |
All Microarray Probes Designed to Gene DMG400030008
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41262_PI426222305 | JHI_St_60k_v1 | DMT400077169 | CACCAAAACAGGACAAGTCTGCTTCTAATTATCAGACTAAAGTTACTGATCCAACAGGCG |
| CUST_41258_PI426222305 | JHI_St_60k_v1 | DMT400077168 | ACTTGGCAGAGAAATTCAAGCCGCGAGATGAAGATAAAGCACTTTCTGAGGTCATTTCAG |
Microarray Signals from CUST_41258_PI426222305
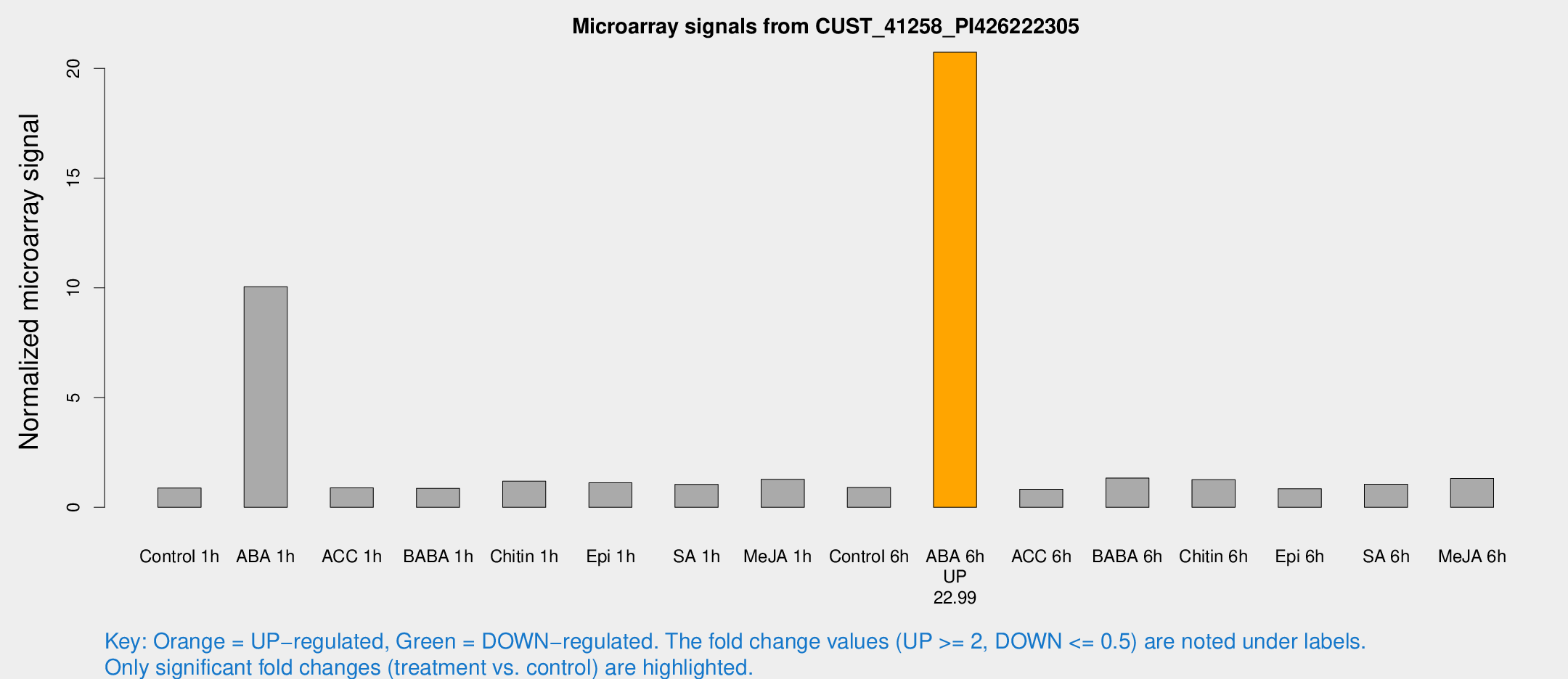
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.93857 | 3.04629 | 0.879437 | 0.460459 |
| ABA 1h | 68.8983 | 21.6928 | 10.0493 | 4.07724 |
| ACC 1h | 6.16784 | 3.63 | 0.885201 | 0.512961 |
| BABA 1h | 5.57806 | 3.22285 | 0.864262 | 0.49869 |
| Chitin 1h | 7.35099 | 3.05217 | 1.19259 | 0.524476 |
| Epi 1h | 6.70955 | 3.00313 | 1.11639 | 0.546953 |
| SA 1h | 7.4826 | 3.1092 | 1.04602 | 0.476672 |
| Me-JA 1h | 7.36091 | 3.15872 | 1.27228 | 0.616328 |
| Control 6h | 6.15399 | 3.11363 | 0.901724 | 0.471682 |
| ABA 6h | 154.528 | 31.1372 | 20.7324 | 4.06247 |
| ACC 6h | 6.46803 | 3.84665 | 0.824079 | 0.478193 |
| BABA 6h | 10.7267 | 3.55211 | 1.33009 | 0.515267 |
| Chitin 6h | 9.51973 | 3.50895 | 1.25867 | 0.523721 |
| Epi 6h | 6.45104 | 3.6698 | 0.845538 | 0.479497 |
| SA 6h | 7.16624 | 3.33921 | 1.0507 | 0.525184 |
| Me-JA 6h | 9.35818 | 3.1718 | 1.31597 | 0.514733 |
Source Transcript PGSC0003DMT400077168 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G52300.1 | +1 | 8e-10 | 59 | 36/110 (33%) | CAP160 protein | chr5:21237205-21239404 FORWARD LENGTH=619 |
