Probe CUST_40943_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40943_PI426222305 | JHI_St_60k_v1 | DMT400079768 | AAATTGAATCACATTTTGCGTTGTTTGTCAAATGATTACAGGGGAAAGCTCTCAATGCCG |
All Microarray Probes Designed to Gene DMG400031063
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40917_PI426222305 | JHI_St_60k_v1 | DMT400079767 | GTCGATTAAGCTTTAAGCCTTTATCGAGAACTTCTCAAATATGTTGCTCTGTTACTTCTA |
| CUST_40943_PI426222305 | JHI_St_60k_v1 | DMT400079768 | AAATTGAATCACATTTTGCGTTGTTTGTCAAATGATTACAGGGGAAAGCTCTCAATGCCG |
Microarray Signals from CUST_40943_PI426222305
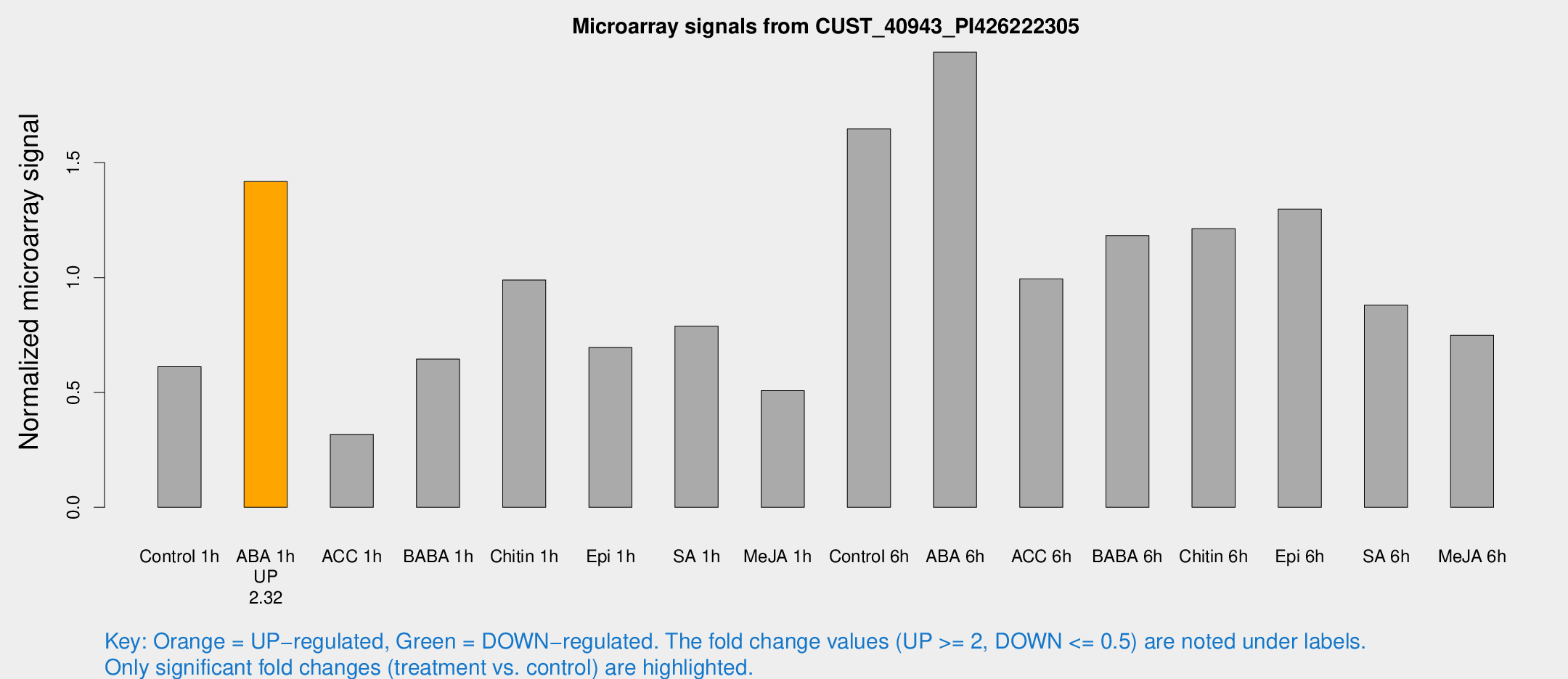
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 21.3715 | 3.98759 | 0.612078 | 0.116414 |
| ABA 1h | 43.812 | 4.44829 | 1.41785 | 0.144079 |
| ACC 1h | 11.5063 | 4.02849 | 0.317902 | 0.116539 |
| BABA 1h | 22.822 | 5.66113 | 0.644944 | 0.171077 |
| Chitin 1h | 31.389 | 4.51605 | 0.989273 | 0.133646 |
| Epi 1h | 24.8663 | 8.52248 | 0.695608 | 0.361433 |
| SA 1h | 32.5887 | 12.2104 | 0.789001 | 0.324185 |
| Me-JA 1h | 17.7191 | 7.76057 | 0.508106 | 0.218567 |
| Control 6h | 59.595 | 12.8198 | 1.64702 | 0.285421 |
| ABA 6h | 80.3211 | 21.6739 | 1.98084 | 0.620025 |
| ACC 6h | 43.7266 | 12.2486 | 0.993861 | 0.269684 |
| BABA 6h | 55.3573 | 19.8994 | 1.18267 | 0.610281 |
| Chitin 6h | 52.9247 | 18.9725 | 1.21314 | 0.607135 |
| Epi 6h | 51.8736 | 8.04388 | 1.29824 | 0.269866 |
| SA 6h | 31.7562 | 7.63308 | 0.880585 | 0.331971 |
| Me-JA 6h | 37.9184 | 16.6543 | 0.748505 | 0.68717 |
Source Transcript PGSC0003DMT400079768 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G58330.1 | +3 | 3e-27 | 113 | 56/73 (77%) | lactate/malate dehydrogenase family protein | chr5:23580010-23582287 REVERSE LENGTH=443 |
