Probe CUST_40917_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40917_PI426222305 | JHI_St_60k_v1 | DMT400079767 | GTCGATTAAGCTTTAAGCCTTTATCGAGAACTTCTCAAATATGTTGCTCTGTTACTTCTA |
All Microarray Probes Designed to Gene DMG400031063
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40917_PI426222305 | JHI_St_60k_v1 | DMT400079767 | GTCGATTAAGCTTTAAGCCTTTATCGAGAACTTCTCAAATATGTTGCTCTGTTACTTCTA |
| CUST_40943_PI426222305 | JHI_St_60k_v1 | DMT400079768 | AAATTGAATCACATTTTGCGTTGTTTGTCAAATGATTACAGGGGAAAGCTCTCAATGCCG |
Microarray Signals from CUST_40917_PI426222305
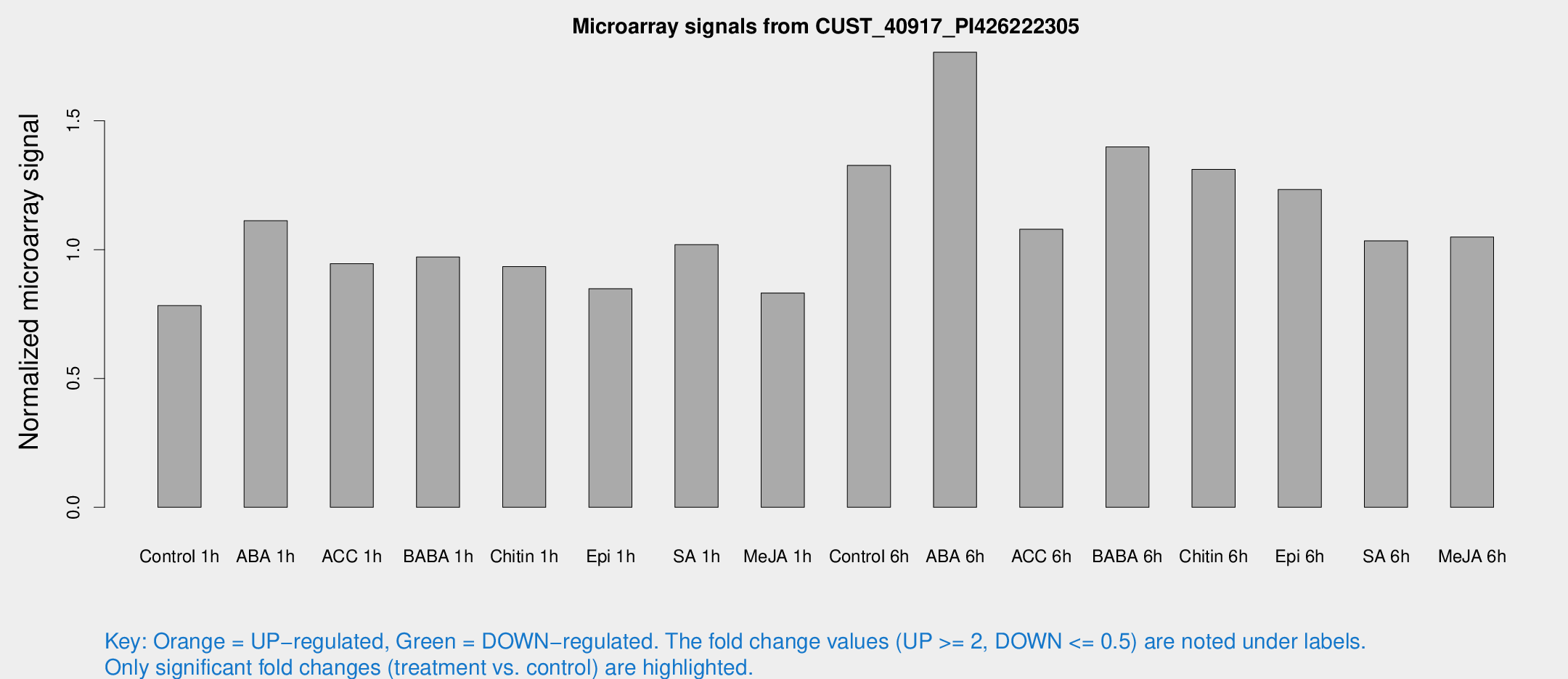
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 240.757 | 40.5835 | 0.782844 | 0.0797778 |
| ABA 1h | 300.466 | 39.0439 | 1.11268 | 0.101377 |
| ACC 1h | 292.564 | 21.3711 | 0.945657 | 0.0559776 |
| BABA 1h | 289.358 | 47.4971 | 0.971319 | 0.0745111 |
| Chitin 1h | 252.263 | 14.9043 | 0.934077 | 0.0560294 |
| Epi 1h | 225.466 | 34.2689 | 0.848373 | 0.118709 |
| SA 1h | 323.453 | 58.827 | 1.01956 | 0.129005 |
| Me-JA 1h | 205.556 | 21.4398 | 0.831907 | 0.049664 |
| Control 6h | 436.948 | 120.013 | 1.32693 | 0.323735 |
| ABA 6h | 594.614 | 125.786 | 1.76637 | 0.3696 |
| ACC 6h | 393.128 | 96.9561 | 1.07908 | 0.117073 |
| BABA 6h | 490.285 | 95.8552 | 1.39914 | 0.263459 |
| Chitin 6h | 446.485 | 115.953 | 1.31154 | 0.31951 |
| Epi 6h | 427.356 | 66.7492 | 1.23375 | 0.263393 |
| SA 6h | 322.511 | 75.0367 | 1.03414 | 0.248356 |
| Me-JA 6h | 365.374 | 118.072 | 1.04897 | 0.403621 |
Source Transcript PGSC0003DMT400079767 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G58330.1 | +3 | 0.0 | 677 | 354/446 (79%) | lactate/malate dehydrogenase family protein | chr5:23580010-23582287 REVERSE LENGTH=443 |
