Probe CUST_40762_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40762_PI426222305 | JHI_St_60k_v1 | DMT400073660 | TTCTTTACTATTGACATCCAACCCTGCCATTCAGTAATCTCTCTGATGTTCTGGTGGAGG |
All Microarray Probes Designed to Gene DMG400028611
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40762_PI426222305 | JHI_St_60k_v1 | DMT400073660 | TTCTTTACTATTGACATCCAACCCTGCCATTCAGTAATCTCTCTGATGTTCTGGTGGAGG |
| CUST_40835_PI426222305 | JHI_St_60k_v1 | DMT400073659 | ATGGTTCTTACTCCAATTGTTTGACATCACACTGATATTTACGCCAAAAGCTCAATTGGA |
Microarray Signals from CUST_40762_PI426222305
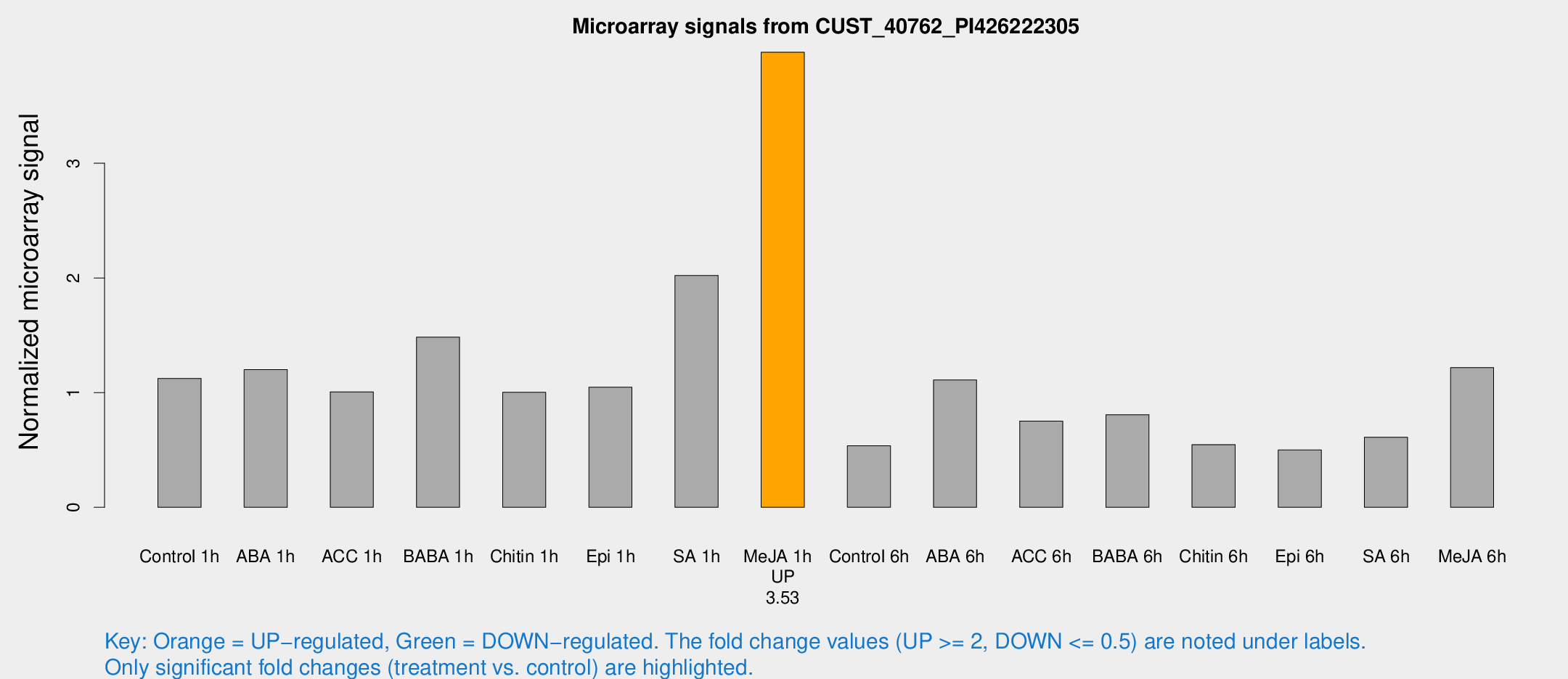
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 182.604 | 29.1118 | 1.12407 | 0.1644 |
| ABA 1h | 173.368 | 30.934 | 1.2009 | 0.12655 |
| ACC 1h | 184.77 | 60.9778 | 1.007 | 0.316144 |
| BABA 1h | 229.88 | 27.7672 | 1.48389 | 0.0892961 |
| Chitin 1h | 146.198 | 23.4908 | 1.00266 | 0.0808951 |
| Epi 1h | 143.862 | 9.03585 | 1.04701 | 0.0657861 |
| SA 1h | 361.193 | 114.525 | 2.02144 | 0.621443 |
| Me-JA 1h | 518.954 | 56.2752 | 3.96828 | 0.230792 |
| Control 6h | 103.474 | 36.3328 | 0.536419 | 0.241699 |
| ABA 6h | 200.302 | 48.2519 | 1.11034 | 0.258435 |
| ACC 6h | 140.402 | 22.0182 | 0.752329 | 0.15482 |
| BABA 6h | 145.213 | 18.087 | 0.807475 | 0.0692479 |
| Chitin 6h | 92.2105 | 6.66189 | 0.546876 | 0.0395293 |
| Epi 6h | 91.6102 | 15.5998 | 0.499853 | 0.0871141 |
| SA 6h | 98.8102 | 16.4132 | 0.611456 | 0.0443639 |
| Me-JA 6h | 192.43 | 11.7238 | 1.21825 | 0.07408 |
Source Transcript PGSC0003DMT400073660 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G05290.1 | +3 | 1e-142 | 418 | 233/302 (77%) | peroxisomal adenine nucleotide carrier 1 | chr3:1506129-1507614 REVERSE LENGTH=322 |
