Probe CUST_40835_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40835_PI426222305 | JHI_St_60k_v1 | DMT400073659 | ATGGTTCTTACTCCAATTGTTTGACATCACACTGATATTTACGCCAAAAGCTCAATTGGA |
All Microarray Probes Designed to Gene DMG400028611
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40762_PI426222305 | JHI_St_60k_v1 | DMT400073660 | TTCTTTACTATTGACATCCAACCCTGCCATTCAGTAATCTCTCTGATGTTCTGGTGGAGG |
| CUST_40835_PI426222305 | JHI_St_60k_v1 | DMT400073659 | ATGGTTCTTACTCCAATTGTTTGACATCACACTGATATTTACGCCAAAAGCTCAATTGGA |
Microarray Signals from CUST_40835_PI426222305
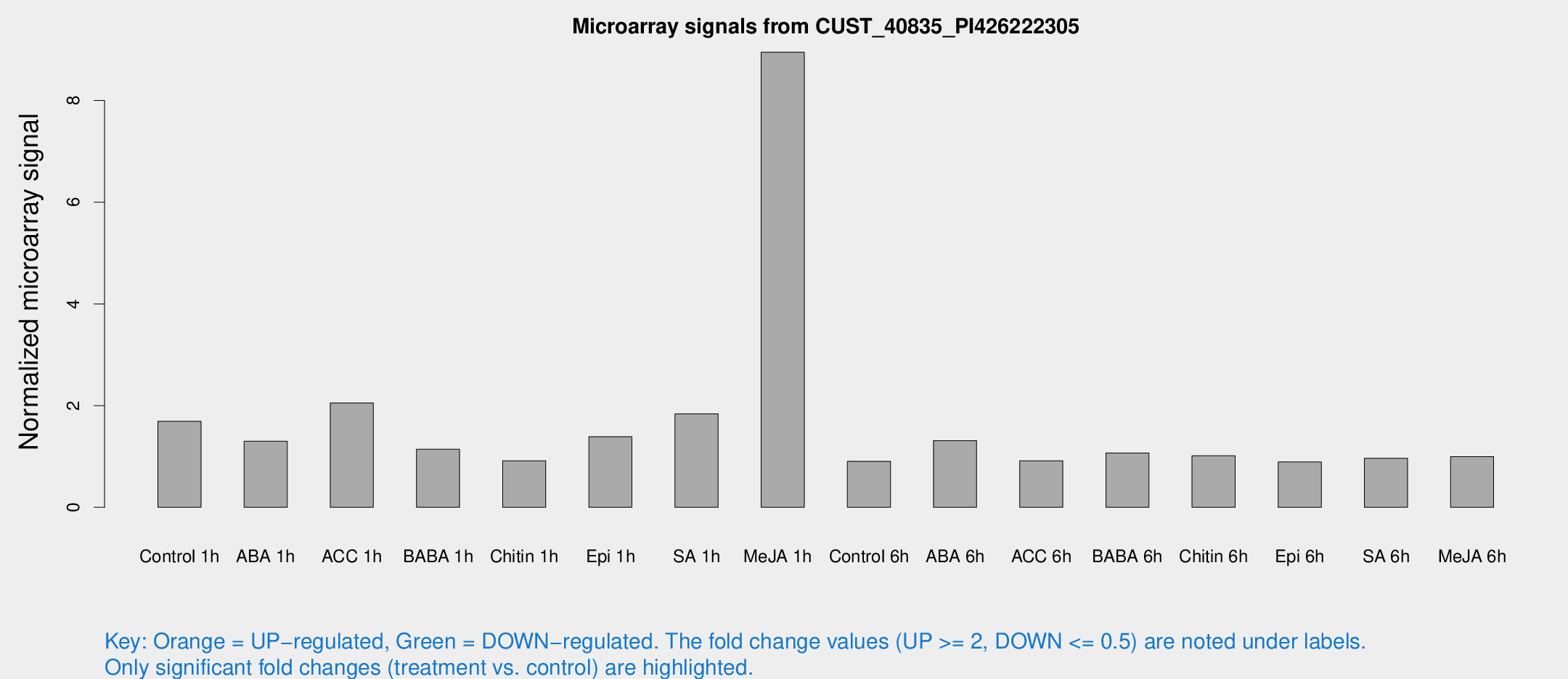
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 13.3888 | 4.42279 | 1.69331 | 0.795208 |
| ABA 1h | 8.23717 | 3.20063 | 1.29819 | 0.551066 |
| ACC 1h | 16.9039 | 5.42666 | 2.05127 | 0.771699 |
| BABA 1h | 7.94167 | 3.54453 | 1.14211 | 0.560572 |
| Chitin 1h | 5.72586 | 3.34436 | 0.915012 | 0.530681 |
| Epi 1h | 9.90989 | 4.35721 | 1.38799 | 0.644446 |
| SA 1h | 15.6821 | 5.54239 | 1.83753 | 0.860545 |
| Me-JA 1h | 51.6895 | 7.77711 | 8.94907 | 0.796023 |
| Control 6h | 6.26913 | 3.64942 | 0.903697 | 0.525365 |
| ABA 6h | 11.5645 | 5.18157 | 1.31047 | 0.60903 |
| ACC 6h | 7.46909 | 4.25711 | 0.914678 | 0.515477 |
| BABA 6h | 8.92082 | 4.02534 | 1.06882 | 0.534002 |
| Chitin 6h | 7.48384 | 4.08786 | 1.01228 | 0.55636 |
| Epi 6h | 7.04835 | 4.14142 | 0.892737 | 0.517693 |
| SA 6h | 6.59748 | 3.82472 | 0.964659 | 0.559101 |
| Me-JA 6h | 6.92663 | 3.62782 | 0.997906 | 0.526056 |
Source Transcript PGSC0003DMT400073659 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G05290.1 | +1 | 8e-82 | 267 | 140/177 (79%) | peroxisomal adenine nucleotide carrier 1 | chr3:1506129-1507614 REVERSE LENGTH=322 |
