Probe CUST_40526_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40526_PI426222305 | JHI_St_60k_v1 | DMT400077598 | CTCAAGATATTGTTGATTCAGCATTTAGAGACCCTGAAGTTAGAAAAGAGAAAGGAAGGA |
All Microarray Probes Designed to Gene DMG400030181
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40526_PI426222305 | JHI_St_60k_v1 | DMT400077598 | CTCAAGATATTGTTGATTCAGCATTTAGAGACCCTGAAGTTAGAAAAGAGAAAGGAAGGA |
| CUST_40502_PI426222305 | JHI_St_60k_v1 | DMT400077597 | CATGGATATGCCATGGAGTGTAGTATATCAATGAAAGGCTATTGAATATGAAACTGTTAG |
Microarray Signals from CUST_40526_PI426222305
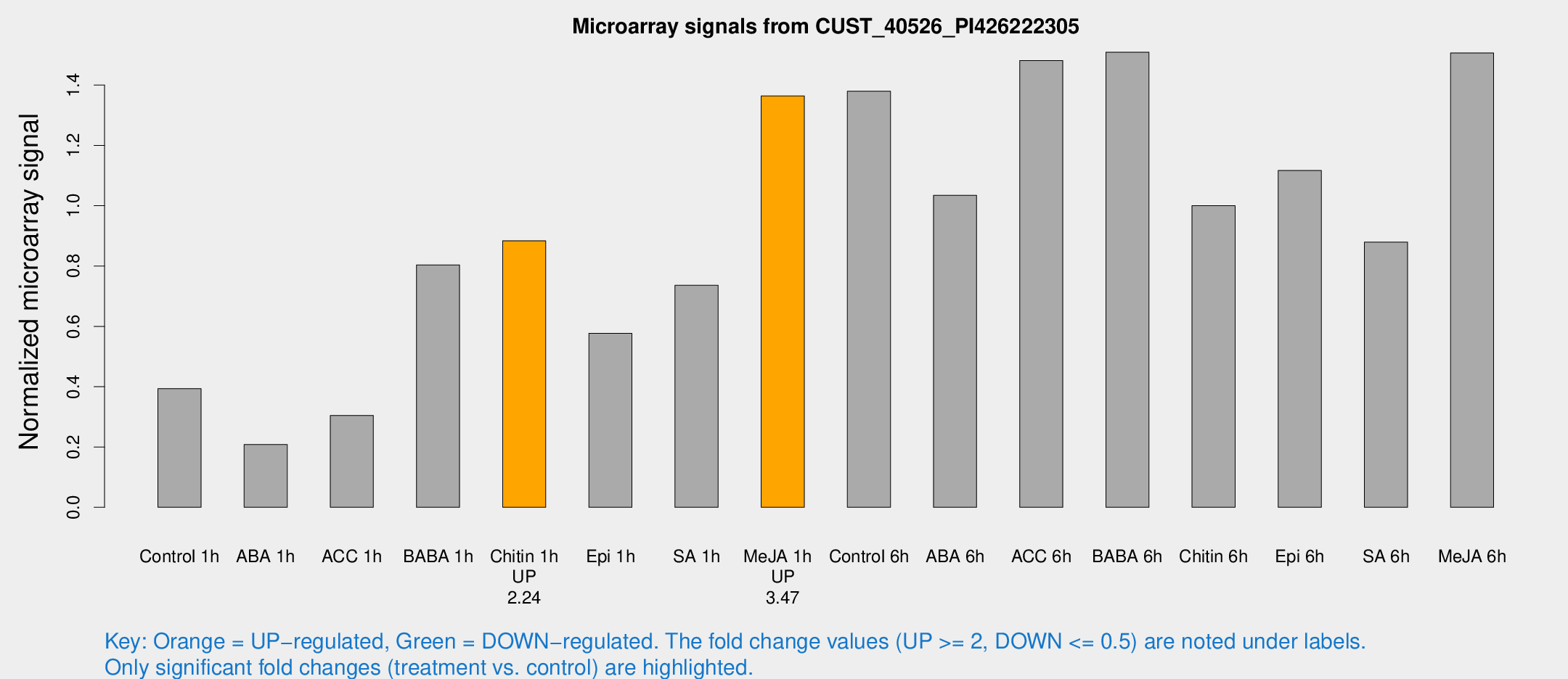
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 955.815 | 149.471 | 0.393648 | 0.0621376 |
| ABA 1h | 489.965 | 177.104 | 0.208036 | 0.0590172 |
| ACC 1h | 866.03 | 306.982 | 0.304655 | 0.117038 |
| BABA 1h | 2037.59 | 577.74 | 0.80372 | 0.204165 |
| Chitin 1h | 1896.5 | 175.126 | 0.883687 | 0.104278 |
| Epi 1h | 1208.1 | 175.148 | 0.57688 | 0.077876 |
| SA 1h | 1839.94 | 317.683 | 0.736222 | 0.0794191 |
| Me-JA 1h | 2710.64 | 471.309 | 1.36403 | 0.115756 |
| Control 6h | 3571.38 | 1029.29 | 1.37994 | 0.332329 |
| ABA 6h | 2787.91 | 692.365 | 1.03466 | 0.232017 |
| ACC 6h | 4115.21 | 635.732 | 1.48149 | 0.0855442 |
| BABA 6h | 4139.63 | 787.813 | 1.50904 | 0.229349 |
| Chitin 6h | 2589.99 | 437.811 | 1.00015 | 0.152857 |
| Epi 6h | 3202.46 | 829.193 | 1.11679 | 0.269212 |
| SA 6h | 2230.32 | 556.721 | 0.879175 | 0.164237 |
| Me-JA 6h | 3753.61 | 840.011 | 1.5065 | 0.294503 |
Source Transcript PGSC0003DMT400077598 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G39420.1 | +2 | 2e-152 | 439 | 201/313 (64%) | alpha/beta-Hydrolases superfamily protein | chr2:16460442-16462872 FORWARD LENGTH=317 |
