Probe CUST_40502_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40502_PI426222305 | JHI_St_60k_v1 | DMT400077597 | CATGGATATGCCATGGAGTGTAGTATATCAATGAAAGGCTATTGAATATGAAACTGTTAG |
All Microarray Probes Designed to Gene DMG400030181
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40526_PI426222305 | JHI_St_60k_v1 | DMT400077598 | CTCAAGATATTGTTGATTCAGCATTTAGAGACCCTGAAGTTAGAAAAGAGAAAGGAAGGA |
| CUST_40502_PI426222305 | JHI_St_60k_v1 | DMT400077597 | CATGGATATGCCATGGAGTGTAGTATATCAATGAAAGGCTATTGAATATGAAACTGTTAG |
Microarray Signals from CUST_40502_PI426222305
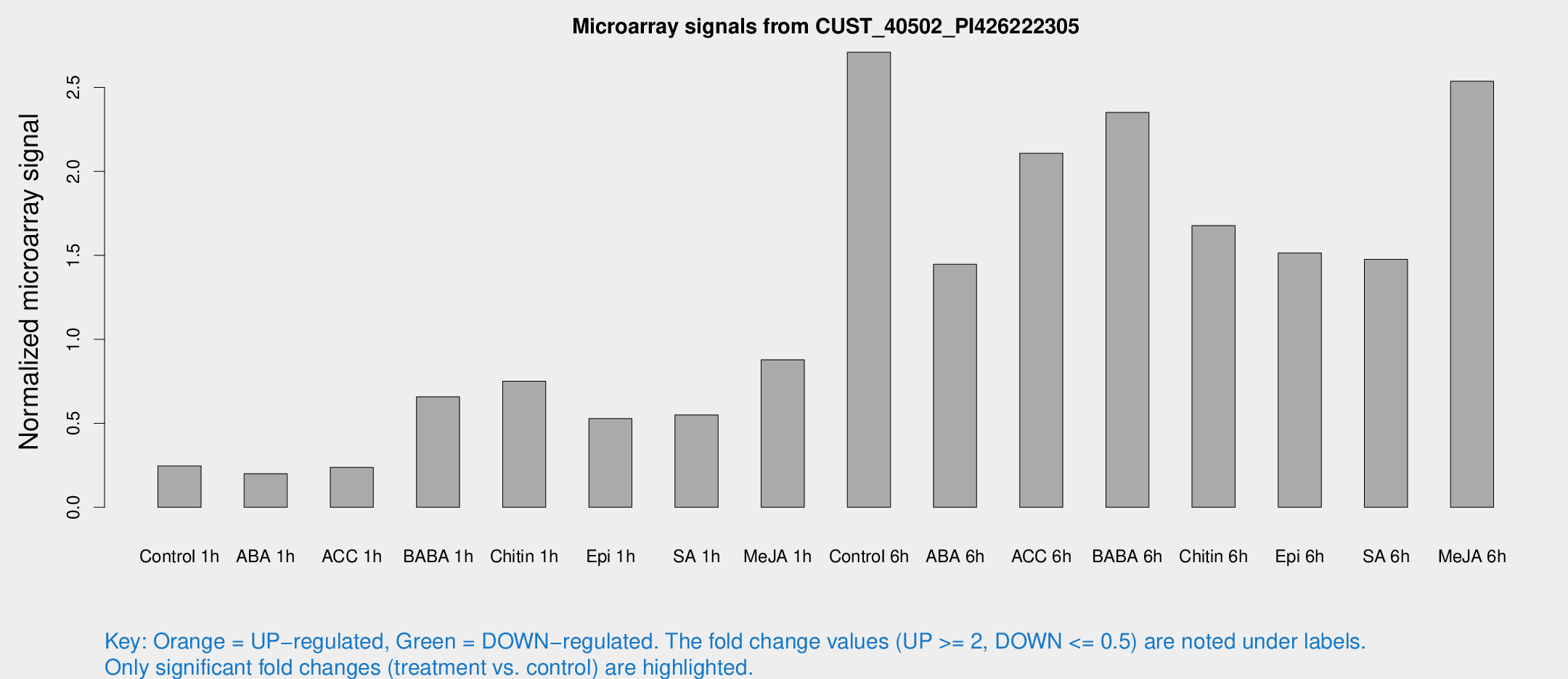
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 13.719 | 3.28957 | 0.247023 | 0.0711449 |
| ABA 1h | 9.81121 | 3.06644 | 0.199773 | 0.0723684 |
| ACC 1h | 17.0008 | 9.01975 | 0.238454 | 0.156281 |
| BABA 1h | 35.9635 | 9.45211 | 0.657866 | 0.134409 |
| Chitin 1h | 36.3407 | 6.12269 | 0.751315 | 0.0808205 |
| Epi 1h | 25.4448 | 6.64658 | 0.528433 | 0.118487 |
| SA 1h | 33.7739 | 12.7591 | 0.550044 | 0.188189 |
| Me-JA 1h | 47.2989 | 21.1863 | 0.878407 | 0.485466 |
| Control 6h | 145.189 | 20.9962 | 2.71004 | 0.169991 |
| ABA 6h | 97.4455 | 44.5202 | 1.4475 | 0.596303 |
| ACC 6h | 129.222 | 18.2426 | 2.1081 | 0.183767 |
| BABA 6h | 164.053 | 71.8937 | 2.35106 | 0.991683 |
| Chitin 6h | 101.522 | 29.3598 | 1.67751 | 0.480294 |
| Epi 6h | 89.9924 | 7.7661 | 1.51413 | 0.107756 |
| SA 6h | 80.9738 | 19.4361 | 1.47714 | 0.41714 |
| Me-JA 6h | 145.467 | 39.1989 | 2.53724 | 0.66453 |
Source Transcript PGSC0003DMT400077597 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G39420.1 | +2 | 8e-93 | 291 | 132/201 (66%) | alpha/beta-Hydrolases superfamily protein | chr2:16460442-16462872 FORWARD LENGTH=317 |
