Probe CUST_40339_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40339_PI426222305 | JHI_St_60k_v1 | DMT400048393 | GAGCCATCTCCAGAAAAGATTGAAATTGCAAATTTATTCATGTTCAACTGTACTATCCCT |
All Microarray Probes Designed to Gene DMG402018799
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40324_PI426222305 | JHI_St_60k_v1 | DMT400048392 | TCCAAACAATTCCGACTCAGTATTGCATATTTTTTGAAAGAACAAGAGAGCCATCTCCAG |
| CUST_40339_PI426222305 | JHI_St_60k_v1 | DMT400048393 | GAGCCATCTCCAGAAAAGATTGAAATTGCAAATTTATTCATGTTCAACTGTACTATCCCT |
Microarray Signals from CUST_40339_PI426222305
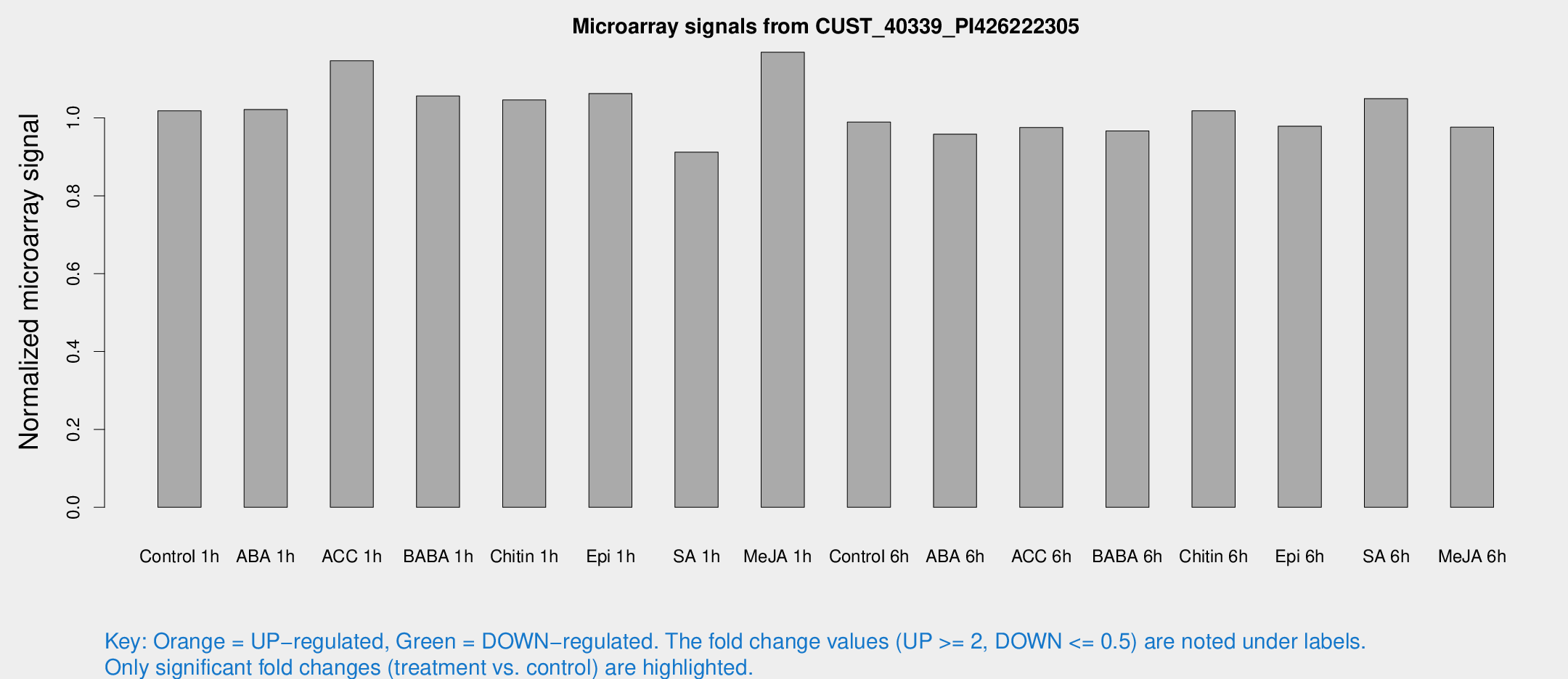
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.15934 | 3.33568 | 1.01815 | 0.552163 |
| ABA 1h | 5.46262 | 3.16496 | 1.02159 | 0.591723 |
| ACC 1h | 7.3279 | 4.01295 | 1.14661 | 0.631153 |
| BABA 1h | 6.17068 | 3.58675 | 1.05623 | 0.612097 |
| Chitin 1h | 5.68786 | 3.31727 | 1.0462 | 0.602837 |
| Epi 1h | 5.56346 | 3.23171 | 1.06269 | 0.614151 |
| SA 1h | 5.67129 | 3.28959 | 0.912107 | 0.528196 |
| Me-JA 1h | 5.81032 | 3.39143 | 1.1687 | 0.676695 |
| Control 6h | 6.00343 | 3.48583 | 0.989549 | 0.573705 |
| ABA 6h | 6.16163 | 3.57104 | 0.958233 | 0.554874 |
| ACC 6h | 6.92374 | 4.11065 | 0.975245 | 0.564735 |
| BABA 6h | 6.55156 | 3.8052 | 0.966347 | 0.560029 |
| Chitin 6h | 6.56007 | 3.80332 | 1.01848 | 0.589725 |
| Epi 6h | 6.76059 | 3.96967 | 0.978872 | 0.566822 |
| SA 6h | 6.2759 | 3.64104 | 1.04968 | 0.608859 |
| Me-JA 6h | 5.87781 | 3.40558 | 0.97627 | 0.565376 |
Source Transcript PGSC0003DMT400048393 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G34040.1 | +1 | 2e-69 | 234 | 114/200 (57%) | Pyridoxal phosphate (PLP)-dependent transferases superfamily protein | chr1:12374433-12376179 FORWARD LENGTH=457 |
