Probe CUST_40324_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40324_PI426222305 | JHI_St_60k_v1 | DMT400048392 | TCCAAACAATTCCGACTCAGTATTGCATATTTTTTGAAAGAACAAGAGAGCCATCTCCAG |
All Microarray Probes Designed to Gene DMG402018799
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40324_PI426222305 | JHI_St_60k_v1 | DMT400048392 | TCCAAACAATTCCGACTCAGTATTGCATATTTTTTGAAAGAACAAGAGAGCCATCTCCAG |
| CUST_40339_PI426222305 | JHI_St_60k_v1 | DMT400048393 | GAGCCATCTCCAGAAAAGATTGAAATTGCAAATTTATTCATGTTCAACTGTACTATCCCT |
Microarray Signals from CUST_40324_PI426222305
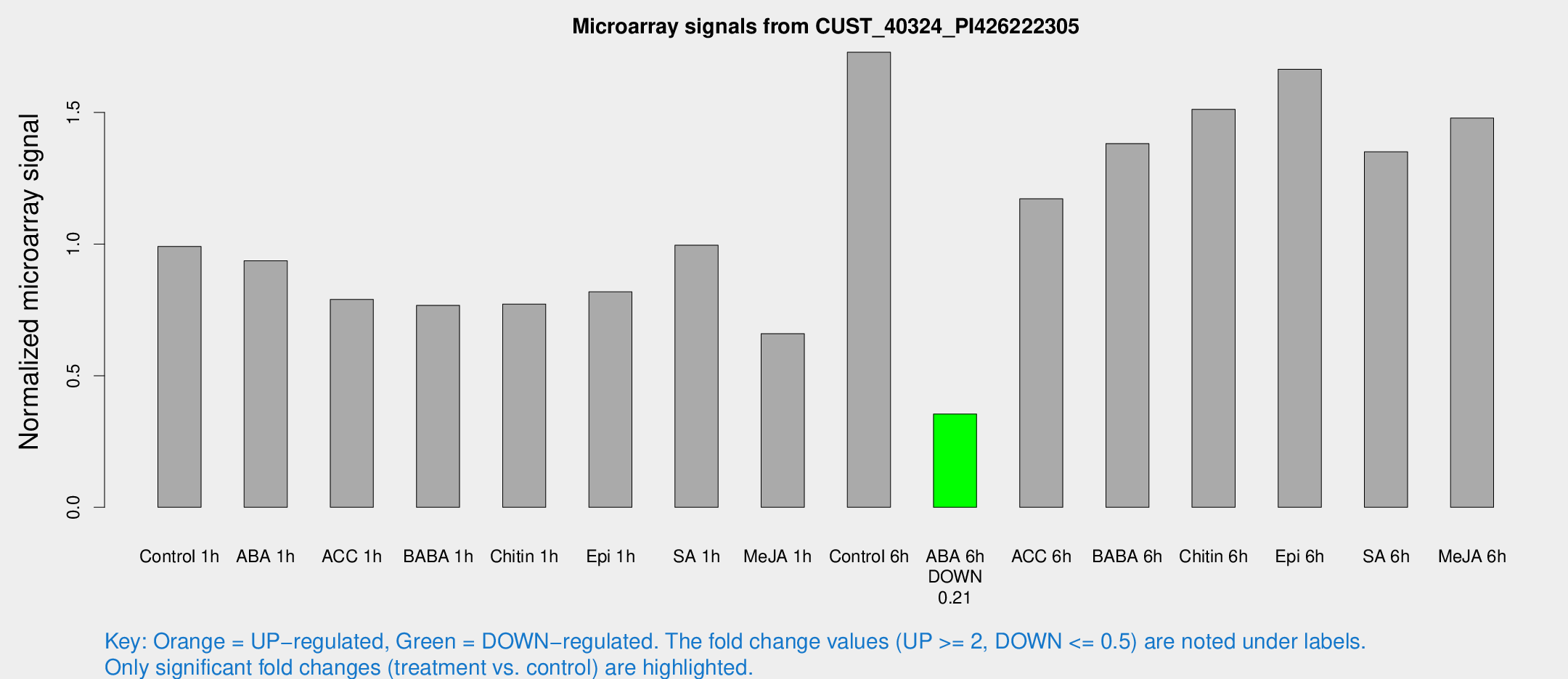
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 119.078 | 27.878 | 0.991078 | 0.161354 |
| ABA 1h | 95.8697 | 11.0648 | 0.936429 | 0.0652702 |
| ACC 1h | 98.9166 | 23.1711 | 0.789566 | 0.155983 |
| BABA 1h | 85.408 | 9.32625 | 0.766989 | 0.053643 |
| Chitin 1h | 80.5326 | 10.8475 | 0.772005 | 0.0540295 |
| Epi 1h | 84.0169 | 15.4245 | 0.818407 | 0.156152 |
| SA 1h | 118.784 | 16.3445 | 0.995539 | 0.128059 |
| Me-JA 1h | 62.4027 | 8.17628 | 0.659226 | 0.0511947 |
| Control 6h | 222.839 | 64.892 | 1.72868 | 0.515403 |
| ABA 6h | 43.0951 | 4.1757 | 0.354455 | 0.0345621 |
| ACC 6h | 162.499 | 40.1618 | 1.17176 | 0.155824 |
| BABA 6h | 179.621 | 23.8288 | 1.3816 | 0.153353 |
| Chitin 6h | 187.642 | 26.2789 | 1.5118 | 0.191019 |
| Epi 6h | 218.465 | 30.0034 | 1.66367 | 0.357839 |
| SA 6h | 154.904 | 19.7155 | 1.35072 | 0.0841159 |
| Me-JA 6h | 172.711 | 27.97 | 1.47851 | 0.12608 |
Source Transcript PGSC0003DMT400048392 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G34060.1 | +2 | 2e-46 | 172 | 82/163 (50%) | Pyridoxal phosphate (PLP)-dependent transferases superfamily protein | chr1:12396561-12398299 REVERSE LENGTH=463 |
