Probe CUST_39995_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39995_PI426222305 | JHI_St_60k_v1 | DMT400030307 | CCTGAGGGAACTTTTGATACCCAAAATGTTTCTTGCGTATTGAACATAAGTAAGAAATTC |
All Microarray Probes Designed to Gene DMG400011597
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39970_PI426222305 | JHI_St_60k_v1 | DMT400030306 | CCTGAGGGAACTTTTGATACCCAAAATGTTTCTTGCGTATTGAACATAAGTAAGAAATTC |
| CUST_39995_PI426222305 | JHI_St_60k_v1 | DMT400030307 | CCTGAGGGAACTTTTGATACCCAAAATGTTTCTTGCGTATTGAACATAAGTAAGAAATTC |
Microarray Signals from CUST_39995_PI426222305
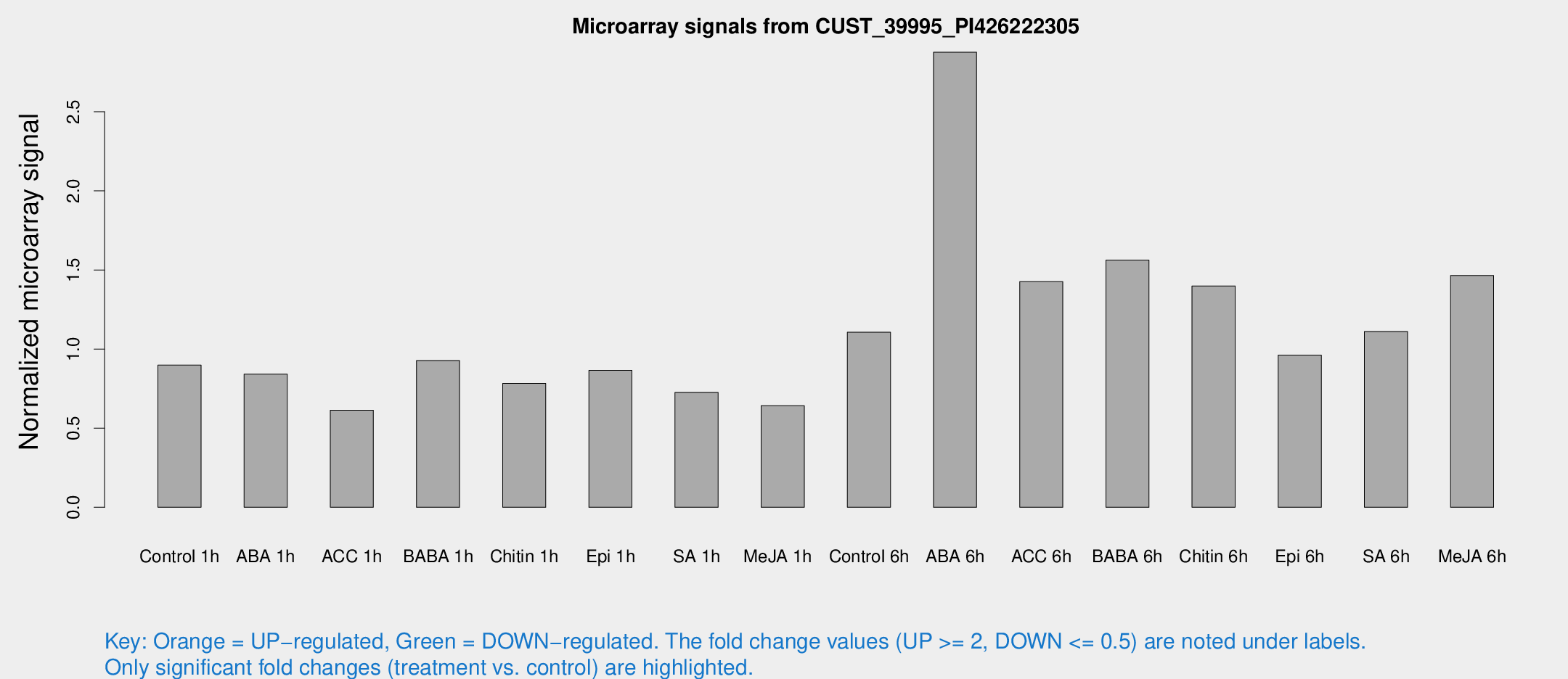
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 214.299 | 16.8505 | 0.898663 | 0.0540632 |
| ABA 1h | 188.787 | 43.7643 | 0.841439 | 0.195728 |
| ACC 1h | 164.743 | 45.055 | 0.613702 | 0.166419 |
| BABA 1h | 219.413 | 38.0957 | 0.927362 | 0.0829306 |
| Chitin 1h | 172.288 | 32.28 | 0.783057 | 0.0915807 |
| Epi 1h | 180.527 | 22.8216 | 0.865349 | 0.109419 |
| SA 1h | 180.239 | 24.0649 | 0.726328 | 0.0607729 |
| Me-JA 1h | 124.916 | 8.86683 | 0.642442 | 0.0416563 |
| Control 6h | 296.898 | 92.784 | 1.10674 | 0.334966 |
| ABA 6h | 733.33 | 81.6946 | 2.876 | 0.166823 |
| ACC 6h | 398.637 | 61.946 | 1.42672 | 0.182382 |
| BABA 6h | 417.819 | 37.6777 | 1.5624 | 0.091643 |
| Chitin 6h | 354.013 | 20.9774 | 1.39916 | 0.0826489 |
| Epi 6h | 259.522 | 24.9588 | 0.961862 | 0.0578953 |
| SA 6h | 265.247 | 36.7921 | 1.1109 | 0.100744 |
| Me-JA 6h | 358.622 | 71.0856 | 1.46483 | 0.168501 |
Source Transcript PGSC0003DMT400030307 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G72660.1 | +2 | 0.0 | 666 | 337/376 (90%) | P-loop containing nucleoside triphosphate hydrolases superfamily protein | chr1:27354161-27356543 FORWARD LENGTH=399 |
