Probe CUST_39970_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39970_PI426222305 | JHI_St_60k_v1 | DMT400030306 | CCTGAGGGAACTTTTGATACCCAAAATGTTTCTTGCGTATTGAACATAAGTAAGAAATTC |
All Microarray Probes Designed to Gene DMG400011597
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39970_PI426222305 | JHI_St_60k_v1 | DMT400030306 | CCTGAGGGAACTTTTGATACCCAAAATGTTTCTTGCGTATTGAACATAAGTAAGAAATTC |
| CUST_39995_PI426222305 | JHI_St_60k_v1 | DMT400030307 | CCTGAGGGAACTTTTGATACCCAAAATGTTTCTTGCGTATTGAACATAAGTAAGAAATTC |
Microarray Signals from CUST_39970_PI426222305
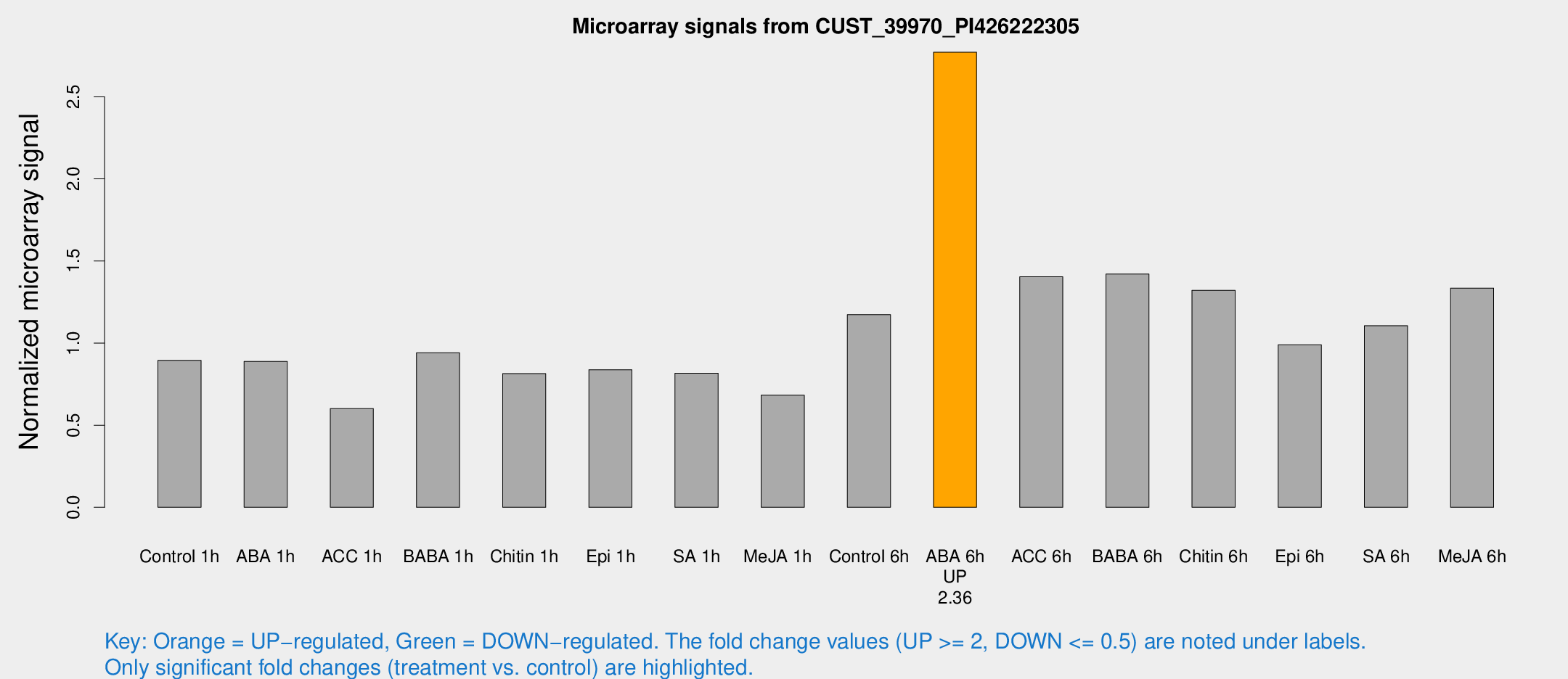
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 211.797 | 12.651 | 0.89511 | 0.0533625 |
| ABA 1h | 200.969 | 51.8099 | 0.888662 | 0.237279 |
| ACC 1h | 164.391 | 49.4836 | 0.601147 | 0.190252 |
| BABA 1h | 221.408 | 37.9384 | 0.940885 | 0.080949 |
| Chitin 1h | 177.146 | 28.595 | 0.814283 | 0.0669051 |
| Epi 1h | 174.526 | 23.3319 | 0.837992 | 0.112966 |
| SA 1h | 204.254 | 33.1605 | 0.81702 | 0.109928 |
| Me-JA 1h | 133.361 | 14.8345 | 0.683334 | 0.0705475 |
| Control 6h | 297.338 | 73.1028 | 1.17301 | 0.23056 |
| ABA 6h | 701.056 | 59.969 | 2.7713 | 0.160618 |
| ACC 6h | 386.831 | 51.7221 | 1.40394 | 0.0823264 |
| BABA 6h | 380.943 | 44.2909 | 1.42032 | 0.120654 |
| Chitin 6h | 334.318 | 27.0157 | 1.32153 | 0.0943185 |
| Epi 6h | 267.985 | 33.1271 | 0.989779 | 0.070249 |
| SA 6h | 264.557 | 42.321 | 1.10626 | 0.145447 |
| Me-JA 6h | 329.526 | 71.8457 | 1.33491 | 0.186169 |
Source Transcript PGSC0003DMT400030306 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G72660.1 | +2 | 0.0 | 708 | 360/399 (90%) | P-loop containing nucleoside triphosphate hydrolases superfamily protein | chr1:27354161-27356543 FORWARD LENGTH=399 |
