Probe CUST_39670_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39670_PI426222305 | JHI_St_60k_v1 | DMT400058739 | GTAAGGCTATAACCTTTTAGTAGTCACTTGACAACAACTTTATAAGTTACACGAGAGCAT |
All Microarray Probes Designed to Gene DMG400022818
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39670_PI426222305 | JHI_St_60k_v1 | DMT400058739 | GTAAGGCTATAACCTTTTAGTAGTCACTTGACAACAACTTTATAAGTTACACGAGAGCAT |
| CUST_39666_PI426222305 | JHI_St_60k_v1 | DMT400058738 | GTAAAGGACTTGAAGATTCTTACATTGATGCTCTTGTGCATAAGCTACAATGTCTATTGT |
Microarray Signals from CUST_39670_PI426222305
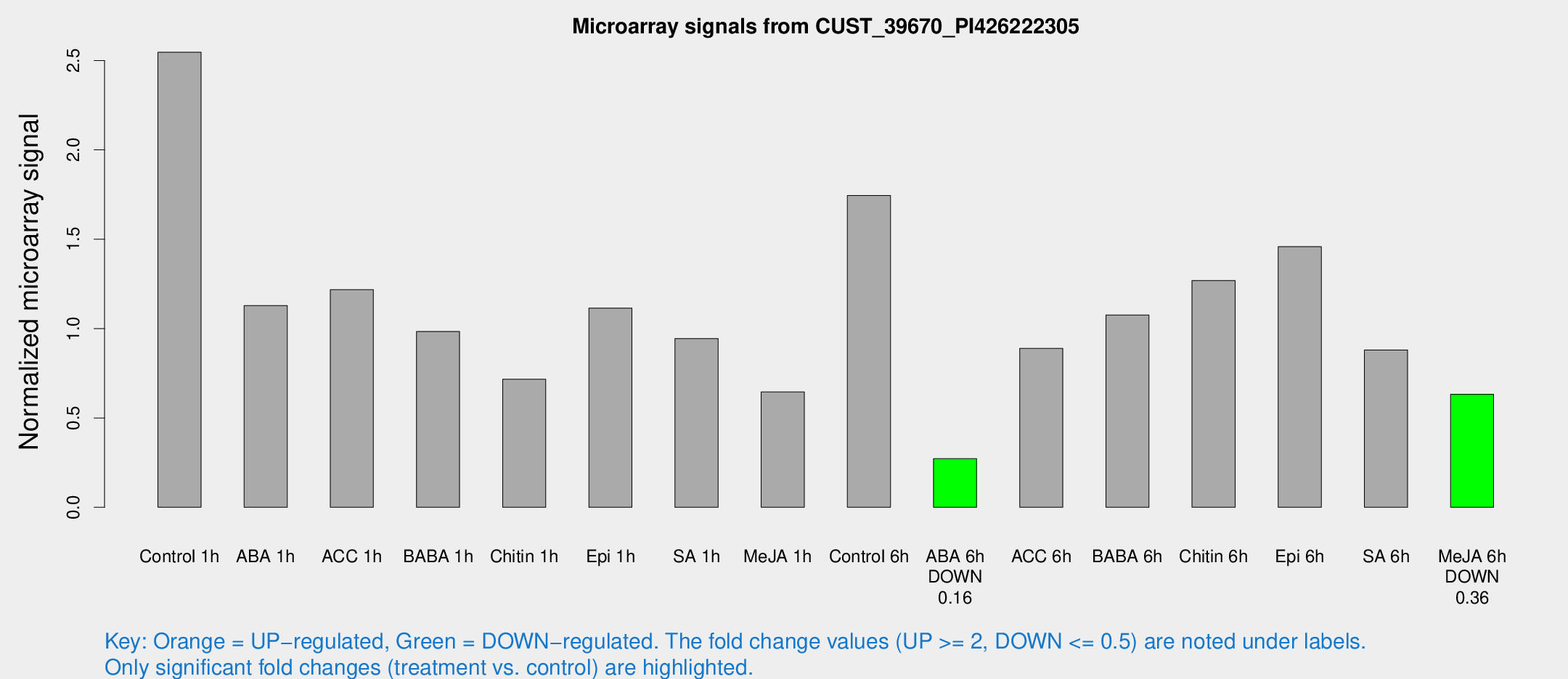
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 270.61 | 62.2396 | 2.54646 | 0.401681 |
| ABA 1h | 102.308 | 12.0214 | 1.12913 | 0.151397 |
| ACC 1h | 131.467 | 25.4025 | 1.21806 | 0.172883 |
| BABA 1h | 109.83 | 38.7204 | 0.983787 | 0.302424 |
| Chitin 1h | 68.4894 | 16.0752 | 0.716275 | 0.107993 |
| Epi 1h | 100.265 | 15.5387 | 1.11503 | 0.17154 |
| SA 1h | 101.762 | 20.4672 | 0.944127 | 0.130666 |
| Me-JA 1h | 53.8895 | 5.80256 | 0.645346 | 0.0619392 |
| Control 6h | 178.756 | 19.2885 | 1.74437 | 0.237474 |
| ABA 6h | 29.6422 | 4.25796 | 0.272204 | 0.0409977 |
| ACC 6h | 105.332 | 15.4466 | 0.888997 | 0.0637038 |
| BABA 6h | 123.748 | 16.079 | 1.07612 | 0.126371 |
| Chitin 6h | 137.053 | 9.10466 | 1.26912 | 0.0842869 |
| Epi 6h | 168.193 | 18.2154 | 1.45829 | 0.275985 |
| SA 6h | 96.9997 | 28.1478 | 0.879782 | 0.194999 |
| Me-JA 6h | 71.6555 | 25.8997 | 0.632146 | 0.174975 |
Source Transcript PGSC0003DMT400058739 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G27570.1 | +1 | 3e-131 | 393 | 201/456 (44%) | UDP-Glycosyltransferase superfamily protein | chr4:13763657-13765018 REVERSE LENGTH=453 |
