Probe CUST_39666_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39666_PI426222305 | JHI_St_60k_v1 | DMT400058738 | GTAAAGGACTTGAAGATTCTTACATTGATGCTCTTGTGCATAAGCTACAATGTCTATTGT |
All Microarray Probes Designed to Gene DMG400022818
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39670_PI426222305 | JHI_St_60k_v1 | DMT400058739 | GTAAGGCTATAACCTTTTAGTAGTCACTTGACAACAACTTTATAAGTTACACGAGAGCAT |
| CUST_39666_PI426222305 | JHI_St_60k_v1 | DMT400058738 | GTAAAGGACTTGAAGATTCTTACATTGATGCTCTTGTGCATAAGCTACAATGTCTATTGT |
Microarray Signals from CUST_39666_PI426222305
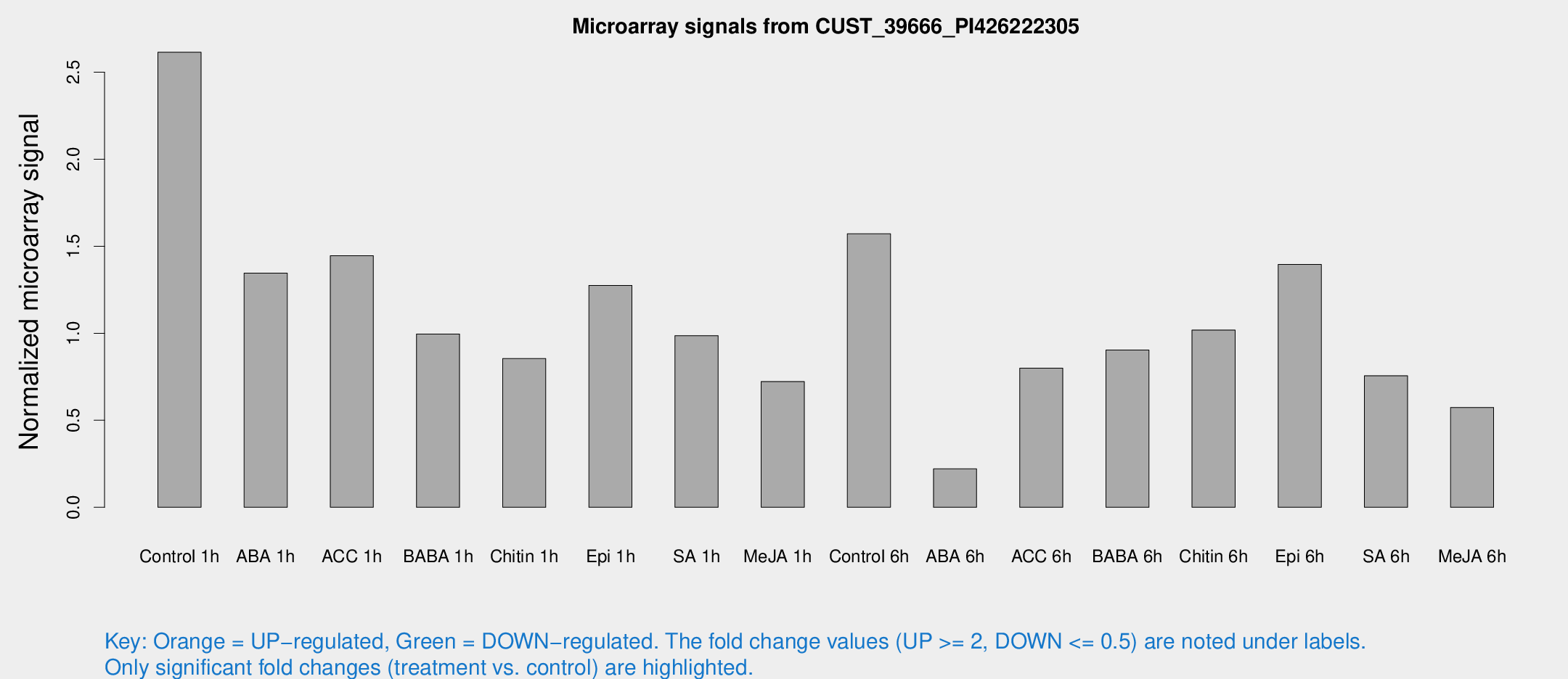
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1084.35 | 251.254 | 2.61471 | 0.453408 |
| ABA 1h | 477.251 | 66.5828 | 1.34505 | 0.146438 |
| ACC 1h | 604.623 | 108.446 | 1.44516 | 0.182582 |
| BABA 1h | 412.076 | 106.757 | 0.995938 | 0.219451 |
| Chitin 1h | 312.448 | 55.7839 | 0.854553 | 0.101163 |
| Epi 1h | 438.588 | 46.8352 | 1.27521 | 0.124979 |
| SA 1h | 407.905 | 61.0264 | 0.986368 | 0.117293 |
| Me-JA 1h | 234.283 | 24.8789 | 0.722841 | 0.0754349 |
| Control 6h | 653.606 | 159.143 | 1.57119 | 0.304785 |
| ABA 6h | 94.536 | 14.9274 | 0.22073 | 0.0432642 |
| ACC 6h | 380.658 | 93.7903 | 0.799622 | 0.0874879 |
| BABA 6h | 409.759 | 75.2286 | 0.903389 | 0.148356 |
| Chitin 6h | 427.357 | 28.2804 | 1.01849 | 0.0593247 |
| Epi 6h | 630.526 | 92.0443 | 1.39526 | 0.320287 |
| SA 6h | 312.173 | 71.7301 | 0.755921 | 0.11069 |
| Me-JA 6h | 253.113 | 91.9998 | 0.573849 | 0.159782 |
Source Transcript PGSC0003DMT400058738 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G27570.1 | +1 | 9e-81 | 254 | 124/259 (48%) | UDP-Glycosyltransferase superfamily protein | chr4:13763657-13765018 REVERSE LENGTH=453 |
