Probe CUST_38640_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38640_PI426222305 | JHI_St_60k_v1 | DMT400076188 | CTTCAAAAAGTTTGCATTTTGTTCACTCCTCTTATAGTCTCATGTGGCTATCTCAAGTAA |
All Microarray Probes Designed to Gene DMG400029624
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38628_PI426222305 | JHI_St_60k_v1 | DMT400076189 | CCTCTCCAATAAATCTTGGAATATATCCCAGTGTATTTGGTACTCATGATTAATGGGAAA |
| CUST_38640_PI426222305 | JHI_St_60k_v1 | DMT400076188 | CTTCAAAAAGTTTGCATTTTGTTCACTCCTCTTATAGTCTCATGTGGCTATCTCAAGTAA |
Microarray Signals from CUST_38640_PI426222305
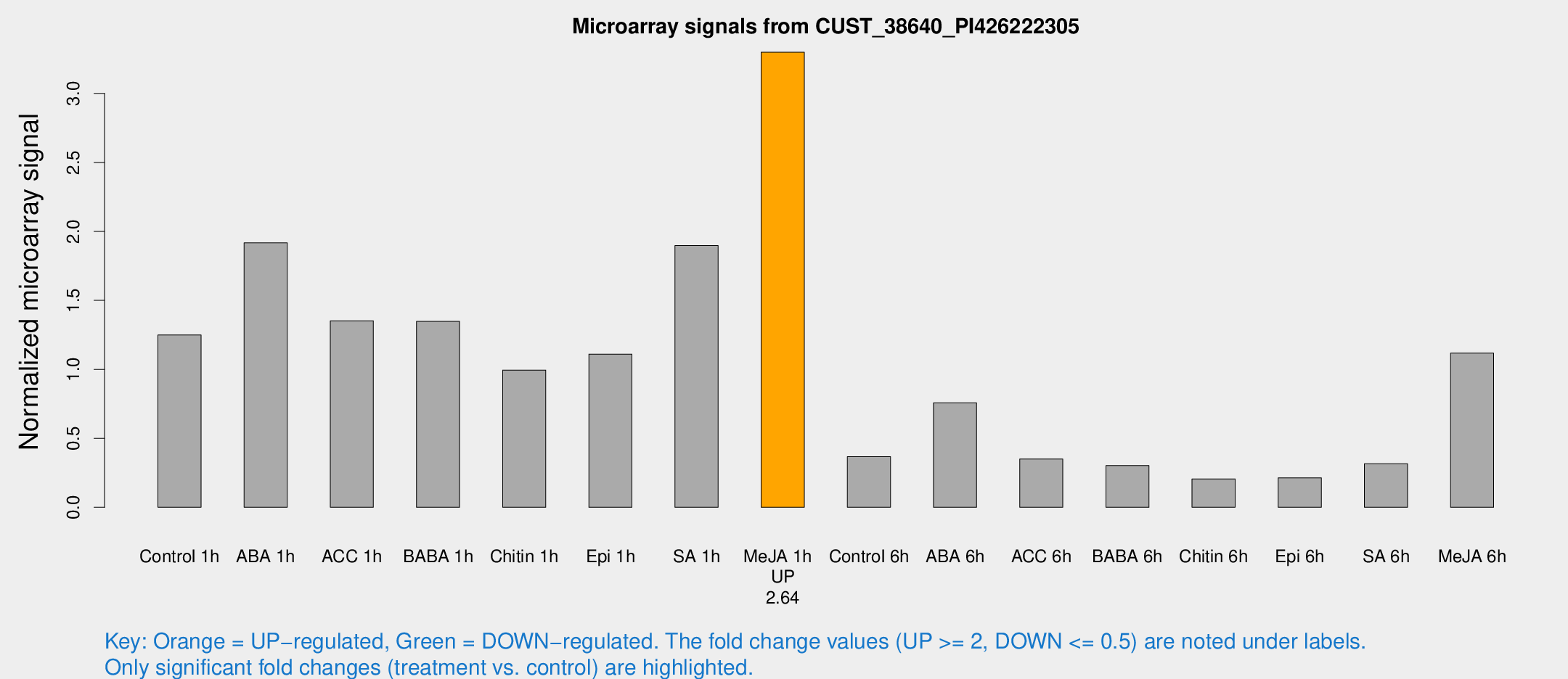
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2076.21 | 537.937 | 1.24958 | 0.232548 |
| ABA 1h | 2700.24 | 364.767 | 1.91657 | 0.128769 |
| ACC 1h | 2358.21 | 679.498 | 1.35201 | 0.329391 |
| BABA 1h | 2160.38 | 478.69 | 1.3479 | 0.212668 |
| Chitin 1h | 1428.45 | 196.038 | 0.994836 | 0.0835488 |
| Epi 1h | 1505.47 | 87.1457 | 1.11048 | 0.0641682 |
| SA 1h | 3269.88 | 900.556 | 1.89748 | 0.561194 |
| Me-JA 1h | 4307.37 | 631.081 | 3.29877 | 0.225441 |
| Control 6h | 682.663 | 231.406 | 0.366811 | 0.145143 |
| ABA 6h | 1440.73 | 519.42 | 0.757145 | 0.257648 |
| ACC 6h | 670.418 | 165.825 | 0.349496 | 0.113607 |
| BABA 6h | 555.808 | 125.685 | 0.302482 | 0.0561374 |
| Chitin 6h | 343.232 | 20.2372 | 0.206085 | 0.012143 |
| Epi 6h | 396.609 | 89.9415 | 0.213681 | 0.0593688 |
| SA 6h | 490.711 | 38.6805 | 0.315278 | 0.0183818 |
| Me-JA 6h | 1903.96 | 559.134 | 1.11725 | 0.489384 |
Source Transcript PGSC0003DMT400076188 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G21950.1 | +1 | 1e-40 | 142 | 76/157 (48%) | S-adenosyl-L-methionine-dependent methyltransferases superfamily protein | chr3:7734396-7736201 FORWARD LENGTH=368 |
