Probe CUST_38628_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38628_PI426222305 | JHI_St_60k_v1 | DMT400076189 | CCTCTCCAATAAATCTTGGAATATATCCCAGTGTATTTGGTACTCATGATTAATGGGAAA |
All Microarray Probes Designed to Gene DMG400029624
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38628_PI426222305 | JHI_St_60k_v1 | DMT400076189 | CCTCTCCAATAAATCTTGGAATATATCCCAGTGTATTTGGTACTCATGATTAATGGGAAA |
| CUST_38640_PI426222305 | JHI_St_60k_v1 | DMT400076188 | CTTCAAAAAGTTTGCATTTTGTTCACTCCTCTTATAGTCTCATGTGGCTATCTCAAGTAA |
Microarray Signals from CUST_38628_PI426222305
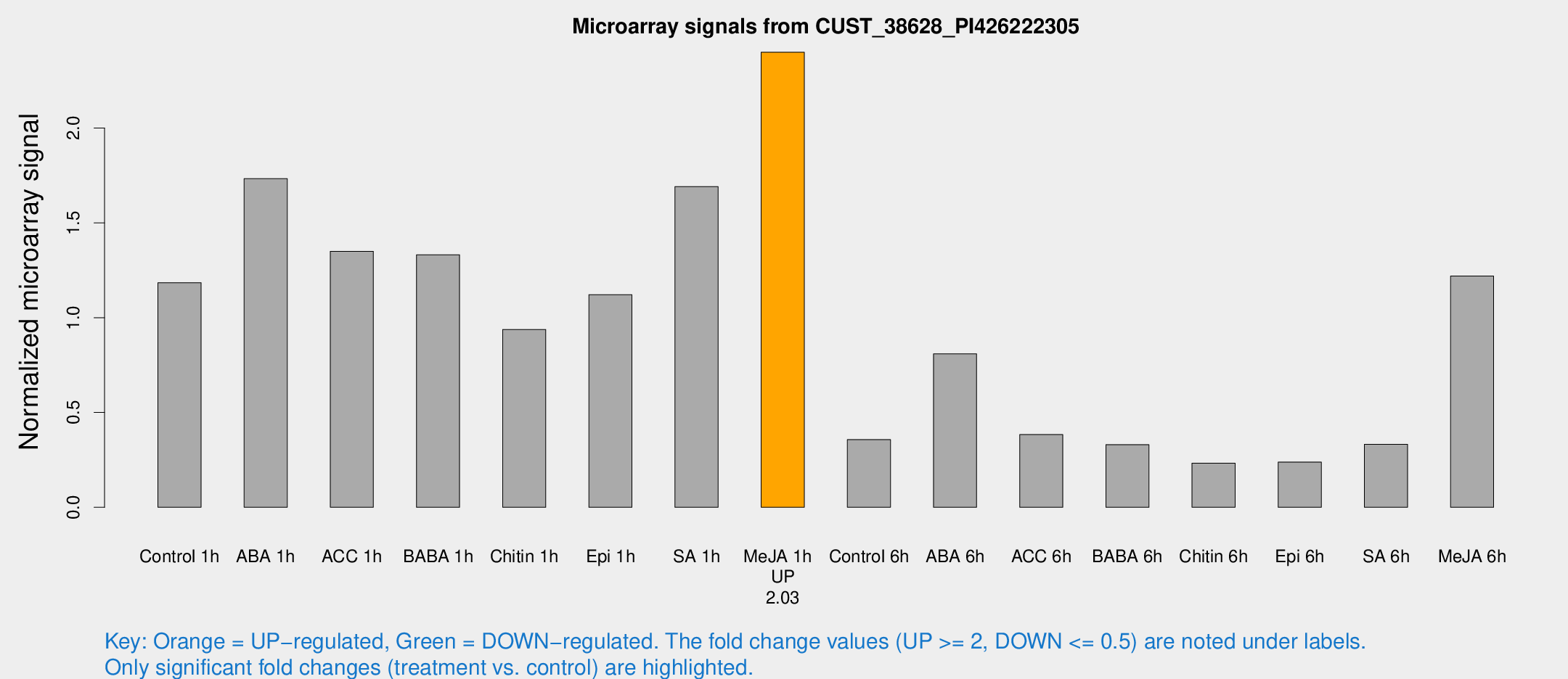
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9405.73 | 1786.62 | 1.18456 | 0.146492 |
| ABA 1h | 11871.5 | 1101.54 | 1.73266 | 0.100036 |
| ACC 1h | 11143.3 | 2480.54 | 1.34998 | 0.253546 |
| BABA 1h | 10127.7 | 1552.75 | 1.33155 | 0.0849275 |
| Chitin 1h | 6538.57 | 619.294 | 0.937604 | 0.054135 |
| Epi 1h | 7477.4 | 432.96 | 1.12128 | 0.0647391 |
| SA 1h | 13949.5 | 2905.64 | 1.69124 | 0.437091 |
| Me-JA 1h | 15290.8 | 1909.04 | 2.39952 | 0.138538 |
| Control 6h | 3166.69 | 1078.27 | 0.357169 | 0.115051 |
| ABA 6h | 7100.45 | 1905.11 | 0.809147 | 0.177438 |
| ACC 6h | 3752.66 | 1195.56 | 0.383755 | 0.161199 |
| BABA 6h | 2999.87 | 721.461 | 0.330277 | 0.0725424 |
| Chitin 6h | 1921.45 | 190.238 | 0.232746 | 0.0264839 |
| Epi 6h | 2121.16 | 338.777 | 0.238564 | 0.051319 |
| SA 6h | 2571.95 | 336.739 | 0.332361 | 0.0191968 |
| Me-JA 6h | 10674.5 | 3838.03 | 1.21909 | 0.656544 |
Source Transcript PGSC0003DMT400076189 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G66430.1 | +2 | 1e-74 | 232 | 127/279 (46%) | S-adenosyl-L-methionine-dependent methyltransferases superfamily protein | chr5:26525410-26526799 REVERSE LENGTH=354 |
