Probe CUST_38470_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38470_PI426222305 | JHI_St_60k_v1 | DMT400050474 | TGATATCAGGTGGCAAAAGTGTATCACAGAATCTGACGACATTCCATACCGCGTCTTTCG |
All Microarray Probes Designed to Gene DMG400019611
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38421_PI426222305 | JHI_St_60k_v1 | DMT400050473 | TTGATGTGCAGCGGCAACTGGAGCTGGATTCATTGCAAGAGACATTACATTTCAGAACTG |
| CUST_38470_PI426222305 | JHI_St_60k_v1 | DMT400050474 | TGATATCAGGTGGCAAAAGTGTATCACAGAATCTGACGACATTCCATACCGCGTCTTTCG |
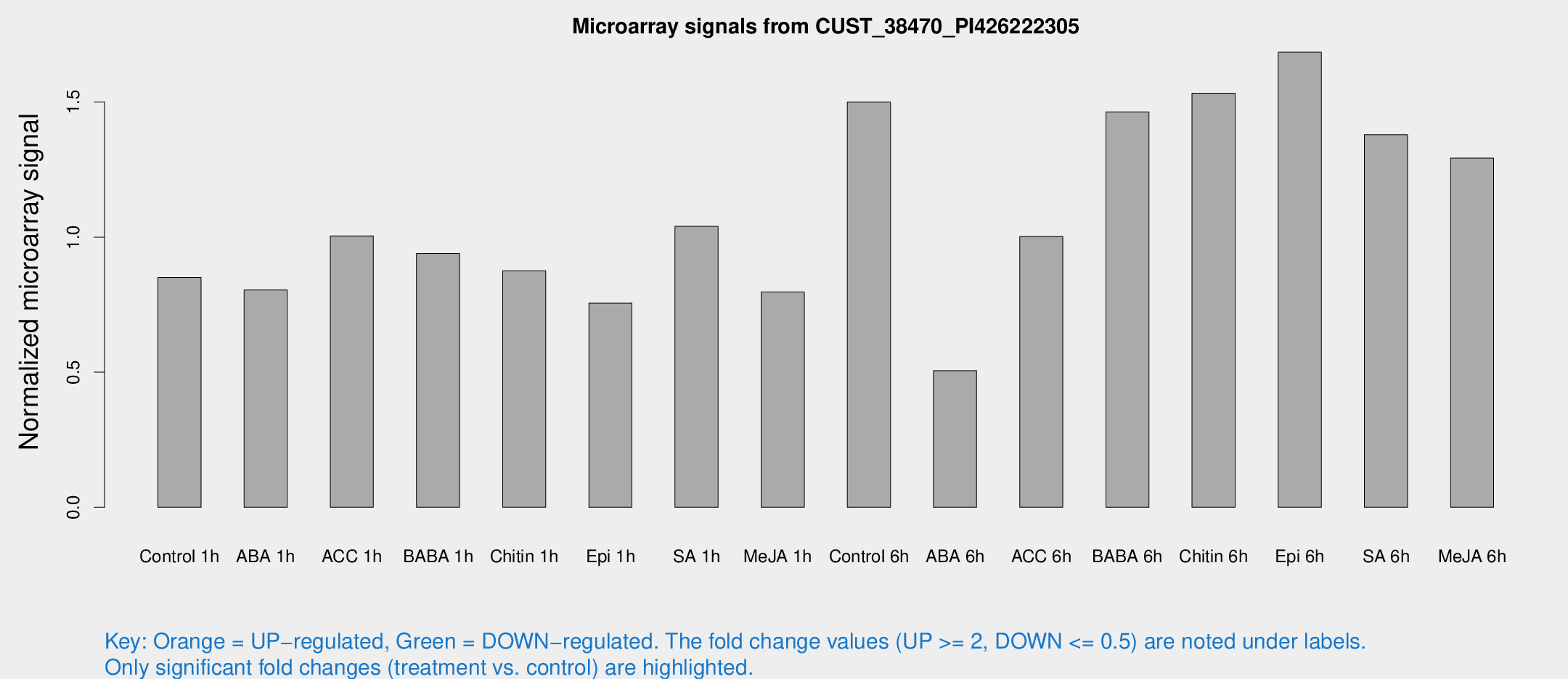
Microarray Signals from CUST_38470_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1127.23 | 416.785 | 0.850803 | 0.266864 |
| ABA 1h | 875.647 | 215.935 | 0.804142 | 0.16656 |
| ACC 1h | 1291.73 | 344.391 | 1.00453 | 0.214622 |
| BABA 1h | 1114.78 | 261.695 | 0.939228 | 0.149632 |
| Chitin 1h | 923.133 | 90.6665 | 0.87552 | 0.0608848 |
| Epi 1h | 766.191 | 76.0149 | 0.755622 | 0.0528333 |
| SA 1h | 1266.9 | 189.683 | 1.03968 | 0.0896812 |
| Me-JA 1h | 773.263 | 114.657 | 0.796762 | 0.0617207 |
| Control 6h | 1947.04 | 570.313 | 1.49983 | 0.418068 |
| ABA 6h | 633.972 | 75.4344 | 0.505834 | 0.0714132 |
| ACC 6h | 1435.77 | 374.076 | 1.00245 | 0.16584 |
| BABA 6h | 1913.61 | 151.715 | 1.46328 | 0.0845405 |
| Chitin 6h | 1902.95 | 135.116 | 1.53286 | 0.120935 |
| Epi 6h | 2257.06 | 359.347 | 1.68437 | 0.299133 |
| SA 6h | 1646.12 | 303.297 | 1.37923 | 0.122523 |
| Me-JA 6h | 1597.5 | 374.735 | 1.2922 | 0.276055 |
Source Transcript PGSC0003DMT400050474 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G49220.1 | +2 | 0.0 | 760 | 374/525 (71%) | Plant invertase/pectin methylesterase inhibitor superfamily | chr3:18249840-18253647 FORWARD LENGTH=598 |
