Probe CUST_38421_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38421_PI426222305 | JHI_St_60k_v1 | DMT400050473 | TTGATGTGCAGCGGCAACTGGAGCTGGATTCATTGCAAGAGACATTACATTTCAGAACTG |
All Microarray Probes Designed to Gene DMG400019611
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38421_PI426222305 | JHI_St_60k_v1 | DMT400050473 | TTGATGTGCAGCGGCAACTGGAGCTGGATTCATTGCAAGAGACATTACATTTCAGAACTG |
| CUST_38470_PI426222305 | JHI_St_60k_v1 | DMT400050474 | TGATATCAGGTGGCAAAAGTGTATCACAGAATCTGACGACATTCCATACCGCGTCTTTCG |
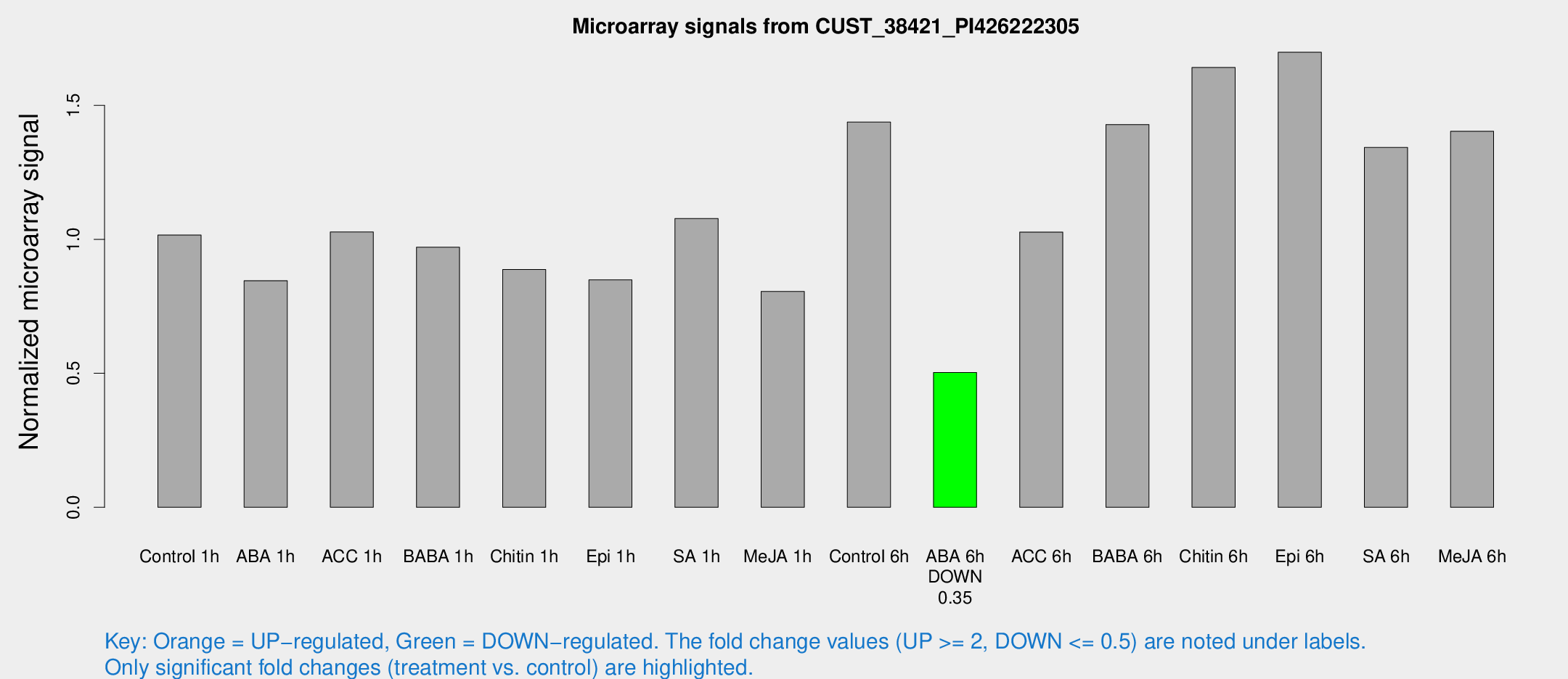
Microarray Signals from CUST_38421_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1007.82 | 300.33 | 1.01573 | 0.226143 |
| ABA 1h | 715.162 | 152.758 | 0.845631 | 0.142438 |
| ACC 1h | 1033.42 | 260.501 | 1.02721 | 0.202161 |
| BABA 1h | 887.527 | 161.645 | 0.970665 | 0.0976282 |
| Chitin 1h | 738.55 | 75.2893 | 0.887296 | 0.0547582 |
| Epi 1h | 678.856 | 67.9962 | 0.848715 | 0.060515 |
| SA 1h | 1036.9 | 161.858 | 1.07773 | 0.100021 |
| Me-JA 1h | 615.457 | 88.896 | 0.805415 | 0.0558553 |
| Control 6h | 1461.91 | 411.665 | 1.43753 | 0.380807 |
| ABA 6h | 493.869 | 44.3091 | 0.503178 | 0.0507922 |
| ACC 6h | 1145.7 | 274.663 | 1.02705 | 0.141319 |
| BABA 6h | 1471.86 | 112.369 | 1.42842 | 0.0941444 |
| Chitin 6h | 1602.08 | 92.7373 | 1.64125 | 0.0981904 |
| Epi 6h | 1792.34 | 278.739 | 1.6982 | 0.268067 |
| SA 6h | 1270.93 | 251.698 | 1.34278 | 0.142063 |
| Me-JA 6h | 1384.02 | 353.674 | 1.40332 | 0.339921 |
Source Transcript PGSC0003DMT400050473 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G49220.1 | +1 | 1e-143 | 422 | 195/231 (84%) | Plant invertase/pectin methylesterase inhibitor superfamily | chr3:18249840-18253647 FORWARD LENGTH=598 |
