Probe CUST_37655_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37655_PI426222305 | JHI_St_60k_v1 | DMT400049495 | TATGCCTAGTCATAACAAGGCAAGCTTGAAATTGTGCTCTATGGCGAGTGGTGGGTGAAA |
All Microarray Probes Designed to Gene DMG400019233
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37655_PI426222305 | JHI_St_60k_v1 | DMT400049495 | TATGCCTAGTCATAACAAGGCAAGCTTGAAATTGTGCTCTATGGCGAGTGGTGGGTGAAA |
| CUST_37639_PI426222305 | JHI_St_60k_v1 | DMT400049494 | TATGCCTAGTCATAACAAGGCAAGCTTGAAATTGTGCTCTATGGCGAGTGGTGGGTGAAA |
Microarray Signals from CUST_37655_PI426222305
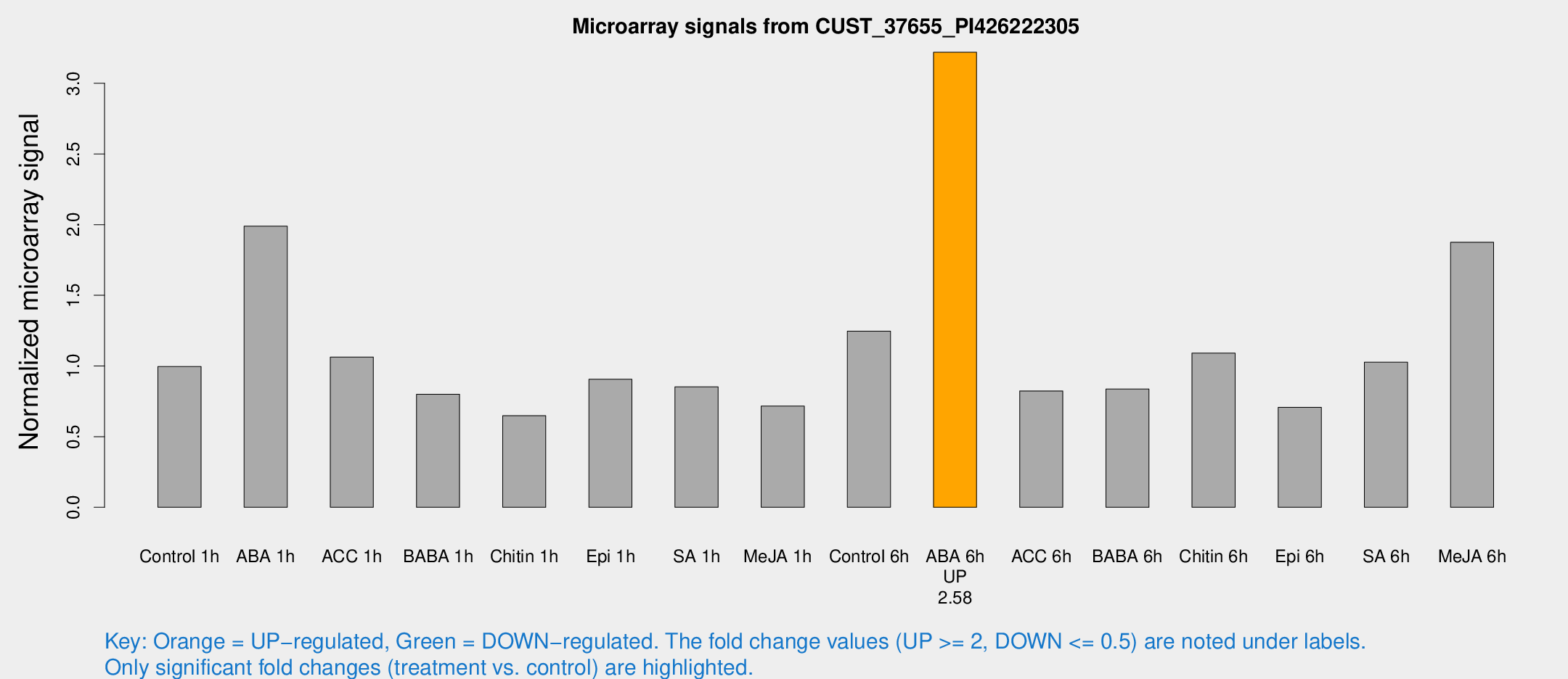
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 34.5705 | 12.5761 | 0.996154 | 0.340609 |
| ABA 1h | 53.8626 | 4.31553 | 1.98911 | 0.159644 |
| ACC 1h | 36.7234 | 9.91981 | 1.06313 | 0.287258 |
| BABA 1h | 24.8375 | 5.33315 | 0.800056 | 0.13114 |
| Chitin 1h | 18.1116 | 3.24632 | 0.648132 | 0.11887 |
| Epi 1h | 26.2766 | 7.00361 | 0.905894 | 0.3068 |
| SA 1h | 26.7691 | 3.49367 | 0.85256 | 0.111465 |
| Me-JA 1h | 20.1507 | 6.12556 | 0.71587 | 0.24381 |
| Control 6h | 39.1545 | 6.77117 | 1.24574 | 0.182077 |
| ABA 6h | 105.877 | 12.5728 | 3.21989 | 0.31146 |
| ACC 6h | 29.7472 | 4.9639 | 0.823283 | 0.169271 |
| BABA 6h | 28.9834 | 3.93716 | 0.836863 | 0.118984 |
| Chitin 6h | 39.0642 | 12.5839 | 1.09092 | 0.376391 |
| Epi 6h | 27.277 | 8.4436 | 0.706866 | 0.323915 |
| SA 6h | 32.6083 | 6.90924 | 1.02677 | 0.140249 |
| Me-JA 6h | 57.6993 | 6.58222 | 1.87532 | 0.151895 |
Source Transcript PGSC0003DMT400049495 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G20660.1 | +2 | 0.0 | 523 | 266/383 (69%) | organic cation/carnitine transporter4 | chr3:7225271-7228510 REVERSE LENGTH=526 |
