Probe CUST_37639_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37639_PI426222305 | JHI_St_60k_v1 | DMT400049494 | TATGCCTAGTCATAACAAGGCAAGCTTGAAATTGTGCTCTATGGCGAGTGGTGGGTGAAA |
All Microarray Probes Designed to Gene DMG400019233
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37655_PI426222305 | JHI_St_60k_v1 | DMT400049495 | TATGCCTAGTCATAACAAGGCAAGCTTGAAATTGTGCTCTATGGCGAGTGGTGGGTGAAA |
| CUST_37639_PI426222305 | JHI_St_60k_v1 | DMT400049494 | TATGCCTAGTCATAACAAGGCAAGCTTGAAATTGTGCTCTATGGCGAGTGGTGGGTGAAA |
Microarray Signals from CUST_37639_PI426222305
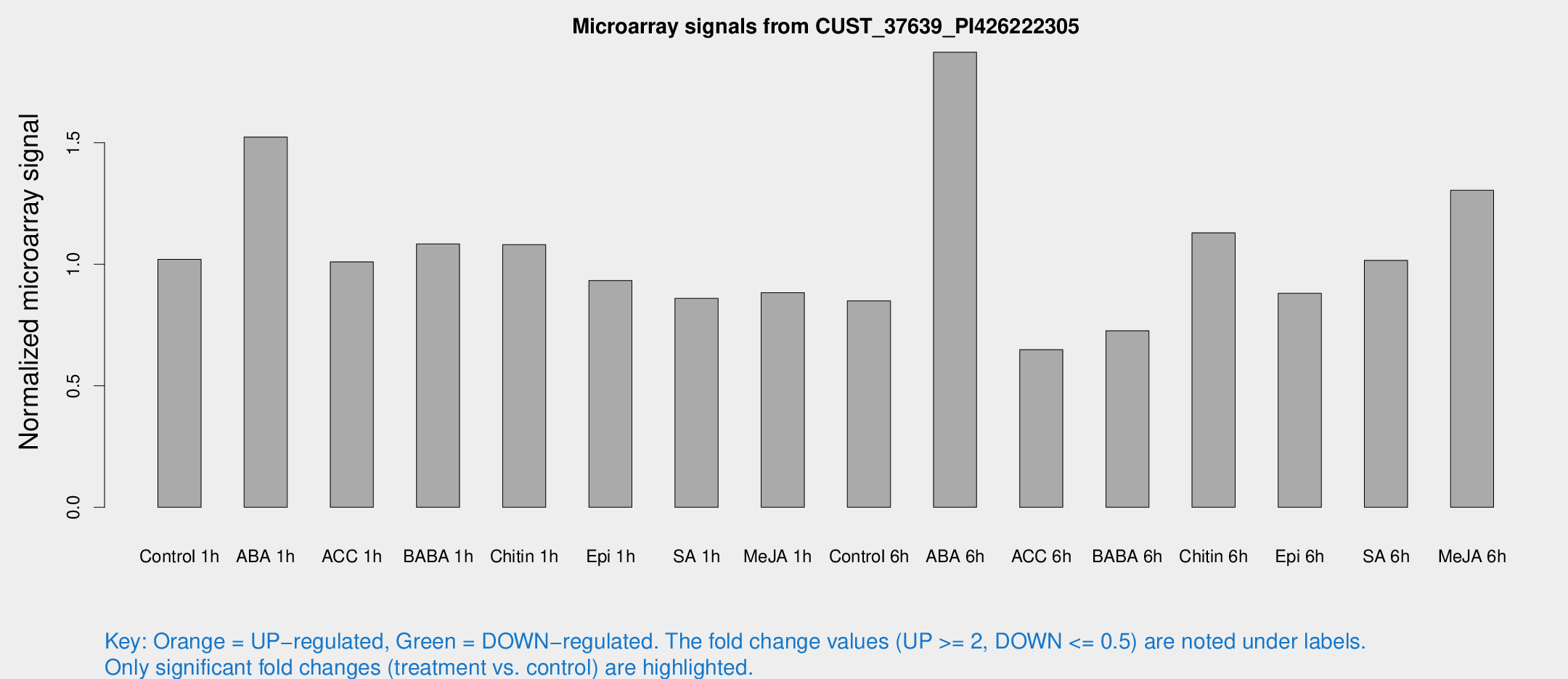
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 67.1164 | 16.9109 | 1.02018 | 0.200042 |
| ABA 1h | 83.7864 | 5.97675 | 1.52358 | 0.108627 |
| ACC 1h | 66.1219 | 11.2718 | 1.01012 | 0.120952 |
| BABA 1h | 64.8584 | 5.58362 | 1.08396 | 0.133063 |
| Chitin 1h | 76.1987 | 37.9562 | 1.08099 | 0.591333 |
| Epi 1h | 50.5548 | 4.85797 | 0.933252 | 0.0891698 |
| SA 1h | 57.1728 | 11.2384 | 0.859859 | 0.154347 |
| Me-JA 1h | 46.17 | 8.15768 | 0.882993 | 0.217579 |
| Control 6h | 53.5327 | 5.92265 | 0.849614 | 0.175861 |
| ABA 6h | 136.922 | 46.2984 | 1.87276 | 0.5147 |
| ACC 6h | 46.9455 | 5.91402 | 0.648563 | 0.0751858 |
| BABA 6h | 52.1205 | 9.45151 | 0.726755 | 0.142039 |
| Chitin 6h | 76.0962 | 10.6766 | 1.12965 | 0.183967 |
| Epi 6h | 62.8272 | 8.34328 | 0.880536 | 0.192209 |
| SA 6h | 67.5731 | 18.0151 | 1.01631 | 0.369596 |
| Me-JA 6h | 83.7437 | 16.9136 | 1.30457 | 0.155005 |
Source Transcript PGSC0003DMT400049494 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G20660.1 | +3 | 0.0 | 679 | 342/517 (66%) | organic cation/carnitine transporter4 | chr3:7225271-7228510 REVERSE LENGTH=526 |
