Probe CUST_37504_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37504_PI426222305 | JHI_St_60k_v1 | DMT400078985 | TTGTGTGTTTTCTTGCATAATTGAATTAATAGCAAAGGGGTGCAACAAGTCAAGCATATC |
All Microarray Probes Designed to Gene DMG401030736
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37445_PI426222305 | JHI_St_60k_v1 | DMT400078986 | TTTTTCATCCACAAGAAGGTCTTCTTTTTGTCTATACACAAGTAACCATGAACATCCTCT |
| CUST_37504_PI426222305 | JHI_St_60k_v1 | DMT400078985 | TTGTGTGTTTTCTTGCATAATTGAATTAATAGCAAAGGGGTGCAACAAGTCAAGCATATC |
Microarray Signals from CUST_37504_PI426222305
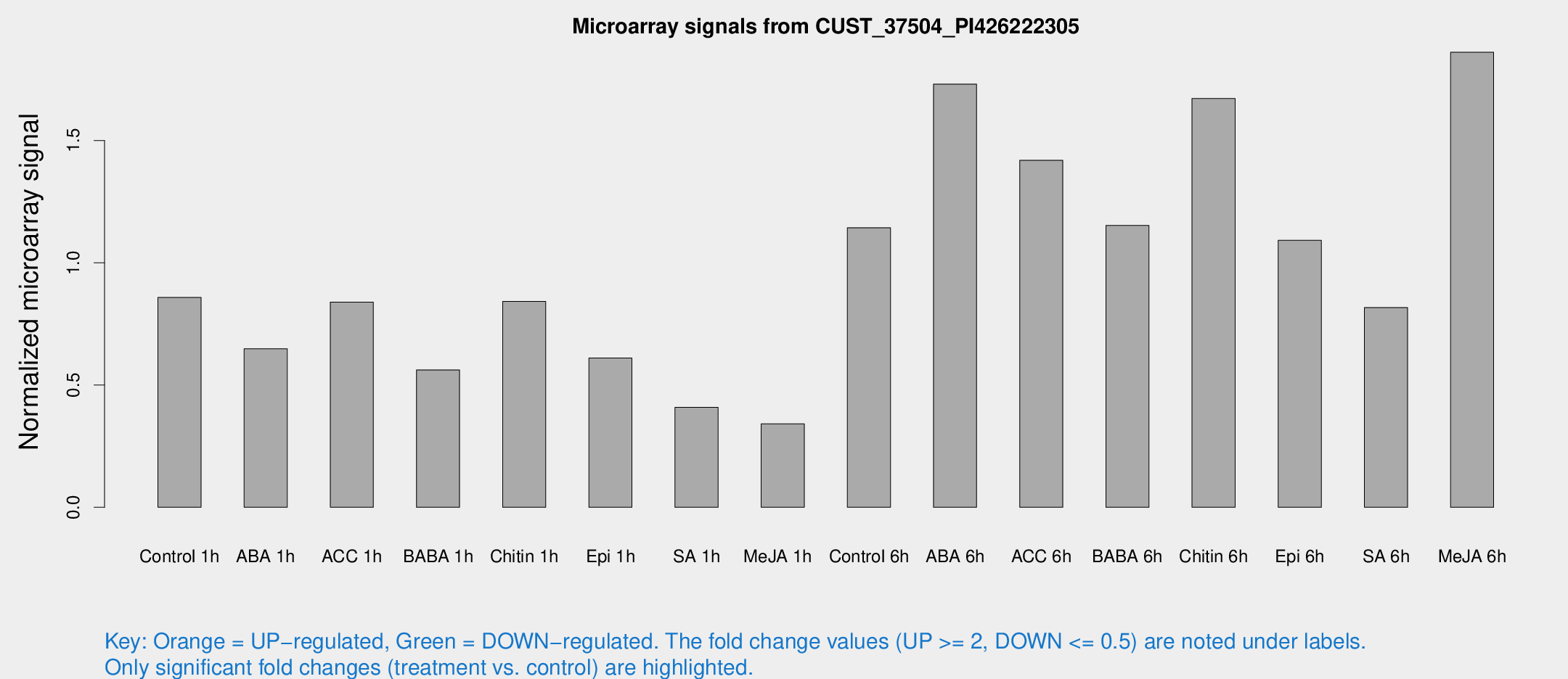
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 31.6306 | 6.92666 | 0.858495 | 0.13633 |
| ABA 1h | 23.6559 | 8.44148 | 0.648397 | 0.275534 |
| ACC 1h | 31.2657 | 5.18426 | 0.839139 | 0.174976 |
| BABA 1h | 25.3896 | 9.97357 | 0.561877 | 0.34881 |
| Chitin 1h | 28.927 | 8.47583 | 0.84189 | 0.20053 |
| Epi 1h | 19.8278 | 4.98525 | 0.61052 | 0.15866 |
| SA 1h | 15.2449 | 3.21982 | 0.408662 | 0.0933536 |
| Me-JA 1h | 10.1623 | 3.22549 | 0.341618 | 0.113041 |
| Control 6h | 52.1202 | 20.1874 | 1.14346 | 0.6457 |
| ABA 6h | 68.5367 | 15.2658 | 1.73121 | 0.328753 |
| ACC 6h | 58.8659 | 9.01117 | 1.41947 | 0.124112 |
| BABA 6h | 48.1293 | 11.0528 | 1.15283 | 0.238462 |
| Chitin 6h | 70.5382 | 20.7171 | 1.67195 | 0.67033 |
| Epi 6h | 44.0263 | 5.32945 | 1.0923 | 0.139851 |
| SA 6h | 32.7289 | 10.0477 | 0.817212 | 0.248226 |
| Me-JA 6h | 68.0642 | 14.2858 | 1.86143 | 0.283096 |
Source Transcript PGSC0003DMT400078985 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G15580.1 | +1 | 7e-25 | 100 | 56/139 (40%) | RING/U-box superfamily protein | chr2:6797687-6798815 FORWARD LENGTH=196 |
