Probe CUST_37445_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37445_PI426222305 | JHI_St_60k_v1 | DMT400078986 | TTTTTCATCCACAAGAAGGTCTTCTTTTTGTCTATACACAAGTAACCATGAACATCCTCT |
All Microarray Probes Designed to Gene DMG401030736
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37445_PI426222305 | JHI_St_60k_v1 | DMT400078986 | TTTTTCATCCACAAGAAGGTCTTCTTTTTGTCTATACACAAGTAACCATGAACATCCTCT |
| CUST_37504_PI426222305 | JHI_St_60k_v1 | DMT400078985 | TTGTGTGTTTTCTTGCATAATTGAATTAATAGCAAAGGGGTGCAACAAGTCAAGCATATC |
Microarray Signals from CUST_37445_PI426222305
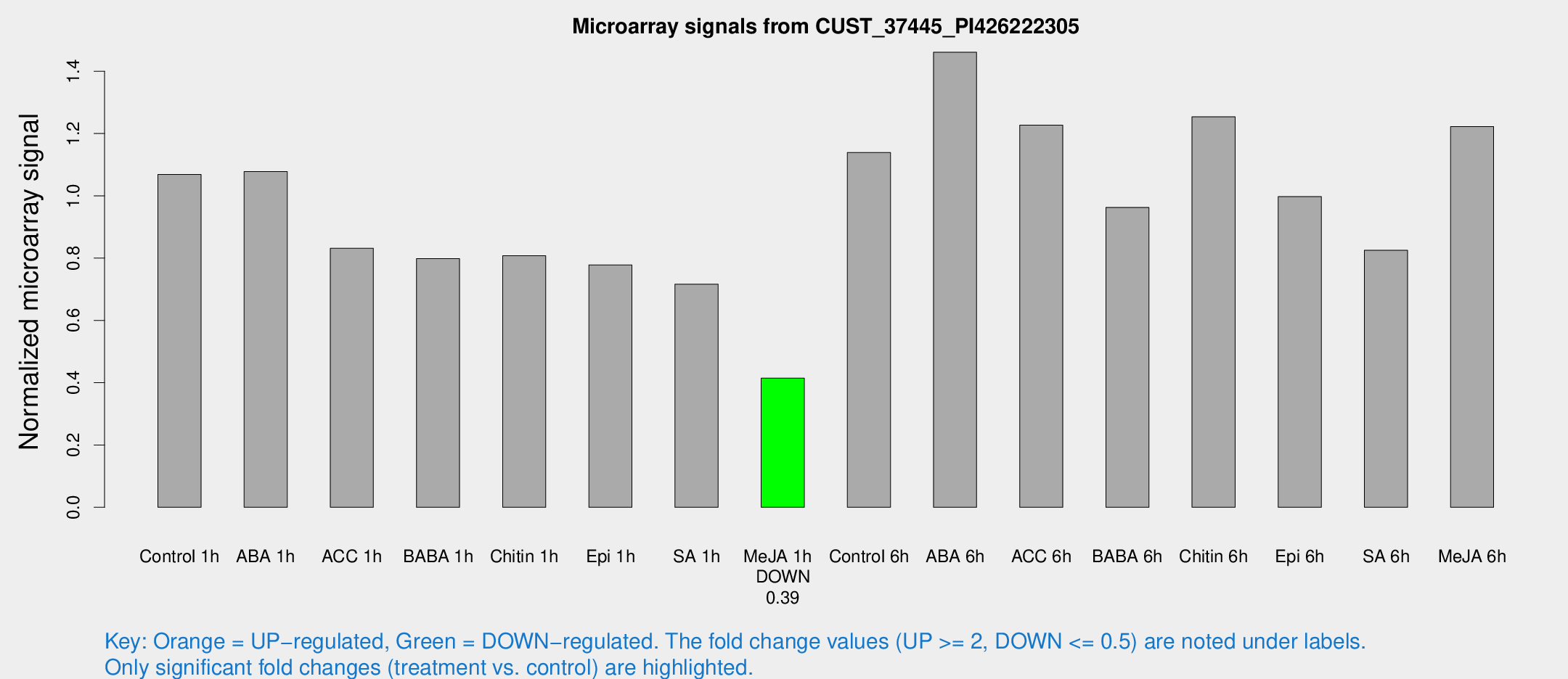
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 455.74 | 28.095 | 1.06849 | 0.0621496 |
| ABA 1h | 409.212 | 39.3517 | 1.07805 | 0.0813607 |
| ACC 1h | 391.743 | 96.1261 | 0.831388 | 0.186107 |
| BABA 1h | 340.307 | 63.4734 | 0.798379 | 0.0837086 |
| Chitin 1h | 320.518 | 65.0979 | 0.807509 | 0.0982567 |
| Epi 1h | 292.081 | 40.3916 | 0.778 | 0.108999 |
| SA 1h | 316.984 | 38.184 | 0.71631 | 0.049656 |
| Me-JA 1h | 146.28 | 17.8789 | 0.41472 | 0.0297214 |
| Control 6h | 515.418 | 120.942 | 1.1391 | 0.205521 |
| ABA 6h | 669.191 | 75.7831 | 1.46098 | 0.101416 |
| ACC 6h | 606.642 | 74.18 | 1.22674 | 0.0712799 |
| BABA 6h | 462.977 | 46.3879 | 0.962734 | 0.0668904 |
| Chitin 6h | 575.936 | 68.6642 | 1.25349 | 0.160187 |
| Epi 6h | 481.635 | 42.9851 | 0.997504 | 0.0937668 |
| SA 6h | 356.698 | 58.4106 | 0.825173 | 0.0678152 |
| Me-JA 6h | 527.279 | 75.4278 | 1.22221 | 0.0854143 |
Source Transcript PGSC0003DMT400078986 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G15580.1 | +2 | 2e-38 | 137 | 88/201 (44%) | RING/U-box superfamily protein | chr2:6797687-6798815 FORWARD LENGTH=196 |
