Probe CUST_36951_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36951_PI426222305 | JHI_St_60k_v1 | DMT400067458 | TTGTATTGAACCAAGGTGGTGCCACCGACATCCATGCTGTAGAAATCTATGAGGAAGAGA |
All Microarray Probes Designed to Gene DMG400026221
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36885_PI426222305 | JHI_St_60k_v1 | DMT400067457 | TATCTTGCTATTGTTGTATTGAACCAAGGTGGTGCCACCGACATCCATGCTGAAGAGAGT |
| CUST_36951_PI426222305 | JHI_St_60k_v1 | DMT400067458 | TTGTATTGAACCAAGGTGGTGCCACCGACATCCATGCTGTAGAAATCTATGAGGAAGAGA |
Microarray Signals from CUST_36951_PI426222305
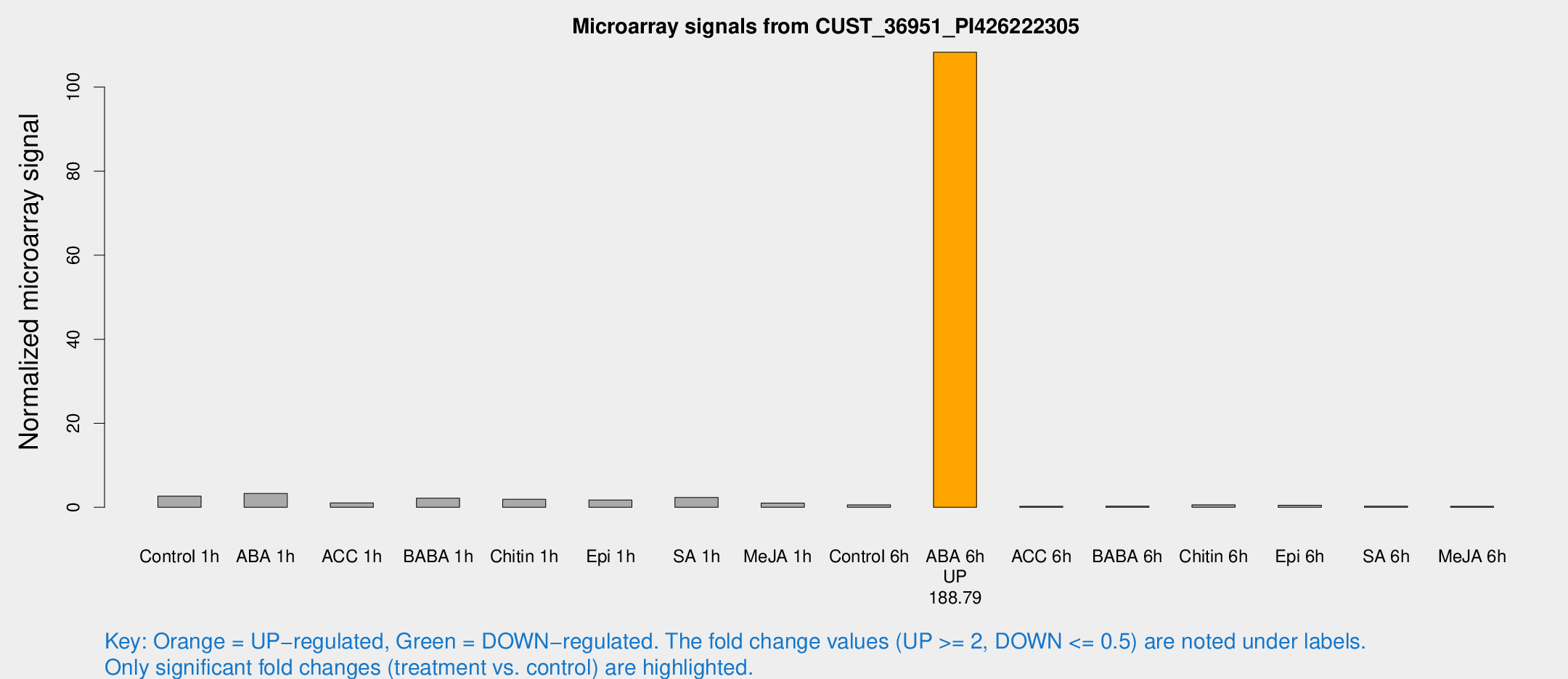
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 156.108 | 68.8561 | 2.65564 | 1.01441 |
| ABA 1h | 161.825 | 54.3317 | 3.29465 | 1.46827 |
| ACC 1h | 106.413 | 60.0194 | 1.05822 | 2.21288 |
| BABA 1h | 104.11 | 7.07999 | 2.18984 | 0.201079 |
| Chitin 1h | 106.503 | 51.113 | 1.90507 | 0.960129 |
| Epi 1h | 86.242 | 27.9202 | 1.73888 | 0.814613 |
| SA 1h | 141.585 | 57.9211 | 2.3358 | 1.40851 |
| Me-JA 1h | 51.7538 | 25.5563 | 1.00385 | 0.676526 |
| Control 6h | 37.5597 | 15.3555 | 0.573685 | 0.360565 |
| ABA 6h | 5736.87 | 614.272 | 108.308 | 6.25358 |
| ACC 6h | 14.8051 | 4.52234 | 0.24579 | 0.100271 |
| BABA 6h | 18.36 | 5.67875 | 0.290167 | 0.138876 |
| Chitin 6h | 46.7124 | 24.7579 | 0.595127 | 0.668144 |
| Epi 6h | 29.5759 | 6.32358 | 0.503424 | 0.146472 |
| SA 6h | 14.1206 | 4.57157 | 0.259112 | 0.0956639 |
| Me-JA 6h | 16.5378 | 10.2787 | 0.230805 | 0.151224 |
Source Transcript PGSC0003DMT400067458 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G17030.1 | +2 | 8e-73 | 229 | 120/238 (50%) | expansin-like B1 | chr4:9581817-9583181 REVERSE LENGTH=250 |
