Probe CUST_36885_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36885_PI426222305 | JHI_St_60k_v1 | DMT400067457 | TATCTTGCTATTGTTGTATTGAACCAAGGTGGTGCCACCGACATCCATGCTGAAGAGAGT |
All Microarray Probes Designed to Gene DMG400026221
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36885_PI426222305 | JHI_St_60k_v1 | DMT400067457 | TATCTTGCTATTGTTGTATTGAACCAAGGTGGTGCCACCGACATCCATGCTGAAGAGAGT |
| CUST_36951_PI426222305 | JHI_St_60k_v1 | DMT400067458 | TTGTATTGAACCAAGGTGGTGCCACCGACATCCATGCTGTAGAAATCTATGAGGAAGAGA |
Microarray Signals from CUST_36885_PI426222305
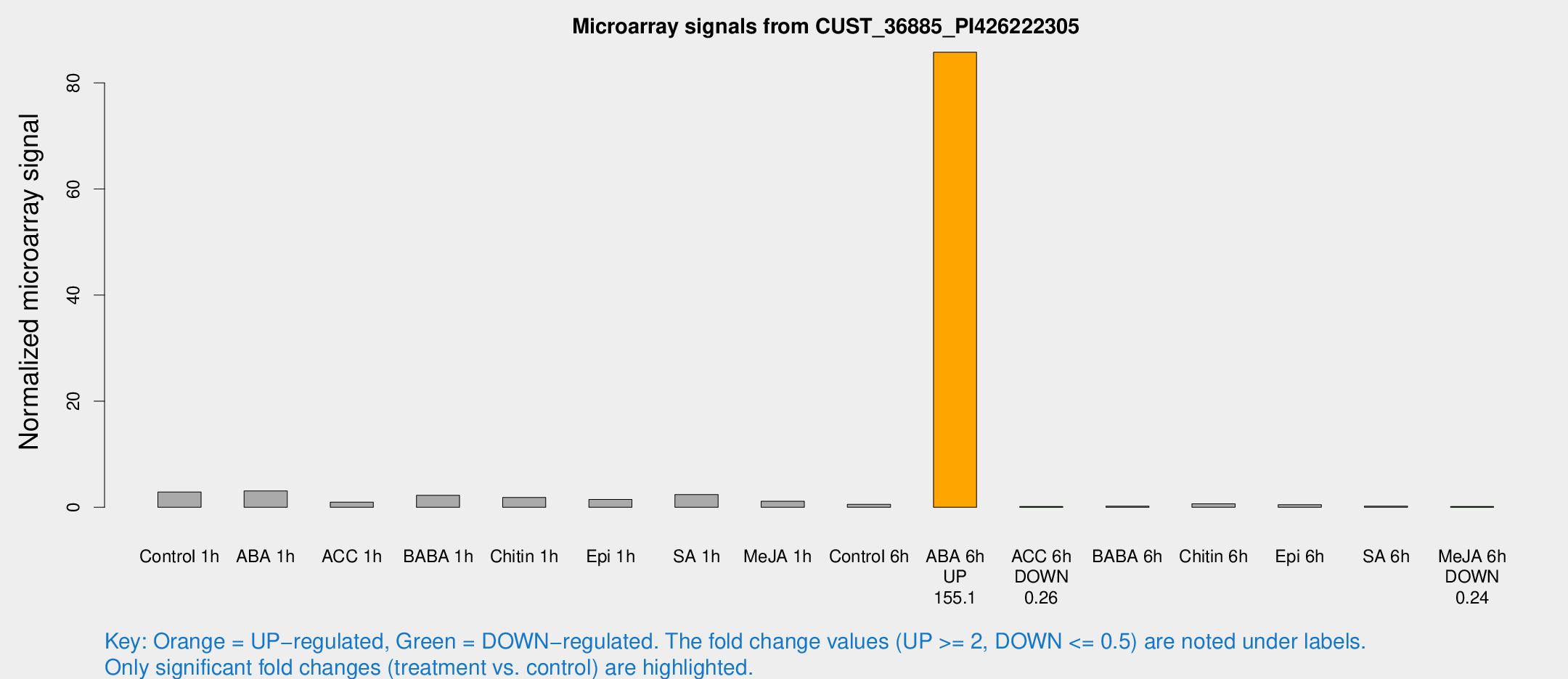
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 198.146 | 71.7447 | 2.88279 | 0.89233 |
| ABA 1h | 185.025 | 59.568 | 3.07238 | 1.32233 |
| ACC 1h | 123.722 | 63.9067 | 0.974971 | 2.38737 |
| BABA 1h | 133.656 | 9.86609 | 2.26373 | 0.391677 |
| Chitin 1h | 141.497 | 64.8717 | 1.84189 | 1.65322 |
| Epi 1h | 103.019 | 40.7067 | 1.47208 | 1.1688 |
| SA 1h | 181.085 | 75.4963 | 2.39835 | 1.48167 |
| Me-JA 1h | 65.5099 | 24.4896 | 1.13908 | 0.570011 |
| Control 6h | 40.2924 | 14.6201 | 0.553049 | 0.224864 |
| ABA 6h | 5585.47 | 410.507 | 85.7761 | 4.95269 |
| ACC 6h | 10.6075 | 4.64842 | 0.14252 | 0.0689248 |
| BABA 6h | 15.4133 | 4.49906 | 0.207743 | 0.0758907 |
| Chitin 6h | 54.4493 | 26.6893 | 0.647734 | 0.434908 |
| Epi 6h | 34.9563 | 8.79034 | 0.474945 | 0.161662 |
| SA 6h | 15.2714 | 4.60148 | 0.226354 | 0.0823419 |
| Me-JA 6h | 8.47235 | 3.82185 | 0.133318 | 0.0639288 |
Source Transcript PGSC0003DMT400067457 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G17030.1 | +1 | 3e-69 | 216 | 118/238 (50%) | expansin-like B1 | chr4:9581817-9583181 REVERSE LENGTH=250 |
