Probe CUST_36917_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36917_PI426222305 | JHI_St_60k_v1 | DMT400067537 | AACCTCGACGTTCTCTATGTAAATACGATATACTTCTCAAGTTGCAAATGAGAAGTTTGA |
All Microarray Probes Designed to Gene DMG400026263
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36894_PI426222305 | JHI_St_60k_v1 | DMT400067538 | GAGGACGTAGCACGACCATTGTCTTATGCTACTTGGTTAAGTACAAGCAGATGACACCAA |
| CUST_36917_PI426222305 | JHI_St_60k_v1 | DMT400067537 | AACCTCGACGTTCTCTATGTAAATACGATATACTTCTCAAGTTGCAAATGAGAAGTTTGA |
Microarray Signals from CUST_36917_PI426222305
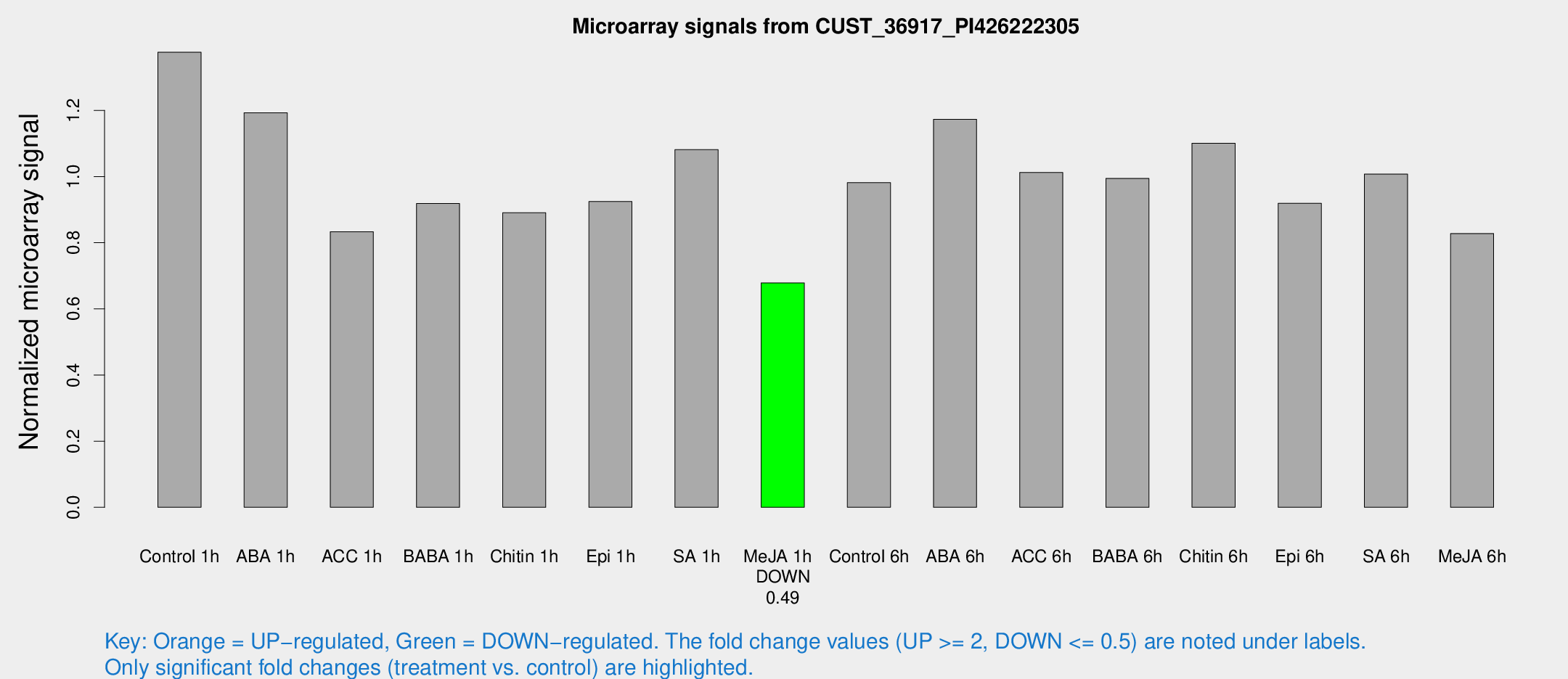
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3060.26 | 177.073 | 1.37589 | 0.0794538 |
| ABA 1h | 2343.05 | 135.351 | 1.193 | 0.0689 |
| ACC 1h | 2023.69 | 456.709 | 0.833266 | 0.160608 |
| BABA 1h | 2083.71 | 472.875 | 0.918418 | 0.141405 |
| Chitin 1h | 1778.88 | 102.863 | 0.890418 | 0.0569249 |
| Epi 1h | 1810.09 | 237.047 | 0.924402 | 0.120936 |
| SA 1h | 2479.77 | 186.239 | 1.08125 | 0.0624454 |
| Me-JA 1h | 1237.79 | 103.262 | 0.678192 | 0.0392076 |
| Control 6h | 2275.71 | 429.81 | 0.981653 | 0.123139 |
| ABA 6h | 2869.41 | 527.253 | 1.17308 | 0.163673 |
| ACC 6h | 2612.45 | 294.626 | 1.01214 | 0.0815621 |
| BABA 6h | 2495.23 | 238.89 | 0.99433 | 0.0645593 |
| Chitin 6h | 2612.74 | 179.887 | 1.10054 | 0.0635679 |
| Epi 6h | 2302.27 | 133.085 | 0.91923 | 0.0531003 |
| SA 6h | 2246.79 | 269.407 | 1.00719 | 0.0581809 |
| Me-JA 6h | 1916.34 | 396.013 | 0.827643 | 0.112627 |
Source Transcript PGSC0003DMT400067537 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G35680.1 | +1 | 4e-14 | 70 | 56/136 (41%) | Phosphotyrosine protein phosphatases superfamily protein | chr2:14997004-14998590 REVERSE LENGTH=337 |
