Probe CUST_36894_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36894_PI426222305 | JHI_St_60k_v1 | DMT400067538 | GAGGACGTAGCACGACCATTGTCTTATGCTACTTGGTTAAGTACAAGCAGATGACACCAA |
All Microarray Probes Designed to Gene DMG400026263
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36894_PI426222305 | JHI_St_60k_v1 | DMT400067538 | GAGGACGTAGCACGACCATTGTCTTATGCTACTTGGTTAAGTACAAGCAGATGACACCAA |
| CUST_36917_PI426222305 | JHI_St_60k_v1 | DMT400067537 | AACCTCGACGTTCTCTATGTAAATACGATATACTTCTCAAGTTGCAAATGAGAAGTTTGA |
Microarray Signals from CUST_36894_PI426222305
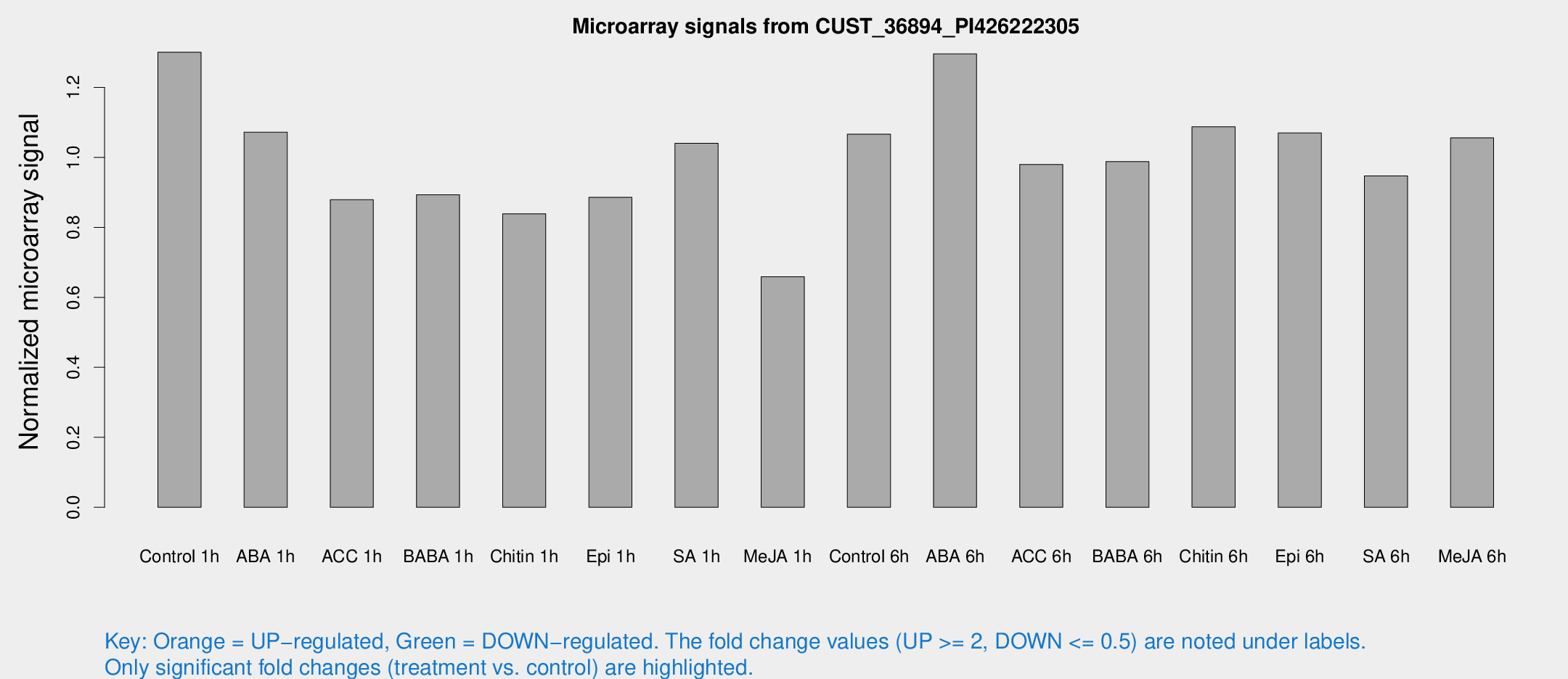
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1520.63 | 215.762 | 1.30038 | 0.101434 |
| ABA 1h | 1097.28 | 109.615 | 1.07176 | 0.0619717 |
| ACC 1h | 1096.6 | 238.552 | 0.878846 | 0.157491 |
| BABA 1h | 1062.13 | 267.512 | 0.893187 | 0.167648 |
| Chitin 1h | 876.205 | 107.004 | 0.838706 | 0.0608321 |
| Epi 1h | 882.333 | 58.4911 | 0.885761 | 0.0512626 |
| SA 1h | 1258.07 | 206.268 | 1.0404 | 0.107759 |
| Me-JA 1h | 625.914 | 81.0953 | 0.658651 | 0.0382272 |
| Control 6h | 1306.38 | 308.186 | 1.06615 | 0.198408 |
| ABA 6h | 1648.29 | 315.301 | 1.29558 | 0.225176 |
| ACC 6h | 1317.79 | 206.042 | 0.979783 | 0.0566614 |
| BABA 6h | 1298.19 | 202.447 | 0.987706 | 0.133905 |
| Chitin 6h | 1333.91 | 106.151 | 1.08723 | 0.0628637 |
| Epi 6h | 1408.03 | 197.479 | 1.06976 | 0.137674 |
| SA 6h | 1191.54 | 339.343 | 0.947066 | 0.222582 |
| Me-JA 6h | 1250.52 | 223.459 | 1.05555 | 0.134822 |
Source Transcript PGSC0003DMT400067538 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G35680.1 | +2 | 7e-100 | 310 | 169/291 (58%) | Phosphotyrosine protein phosphatases superfamily protein | chr2:14997004-14998590 REVERSE LENGTH=337 |
