Probe CUST_36024_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36024_PI426222305 | JHI_St_60k_v1 | DMT400080928 | GGGCGAGTTCTGTAAGGTTAATTAAAGTCGTTACATAATGCCACATCATGTATTATTTAC |
All Microarray Probes Designed to Gene DMG402031520
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36024_PI426222305 | JHI_St_60k_v1 | DMT400080928 | GGGCGAGTTCTGTAAGGTTAATTAAAGTCGTTACATAATGCCACATCATGTATTATTTAC |
| CUST_36021_PI426222305 | JHI_St_60k_v1 | DMT400080927 | GGGCGAGTTCTGTAAGGTTAATTAAAGTCGTTACATAATGCCACATCATGTATTATTTAC |
Microarray Signals from CUST_36024_PI426222305
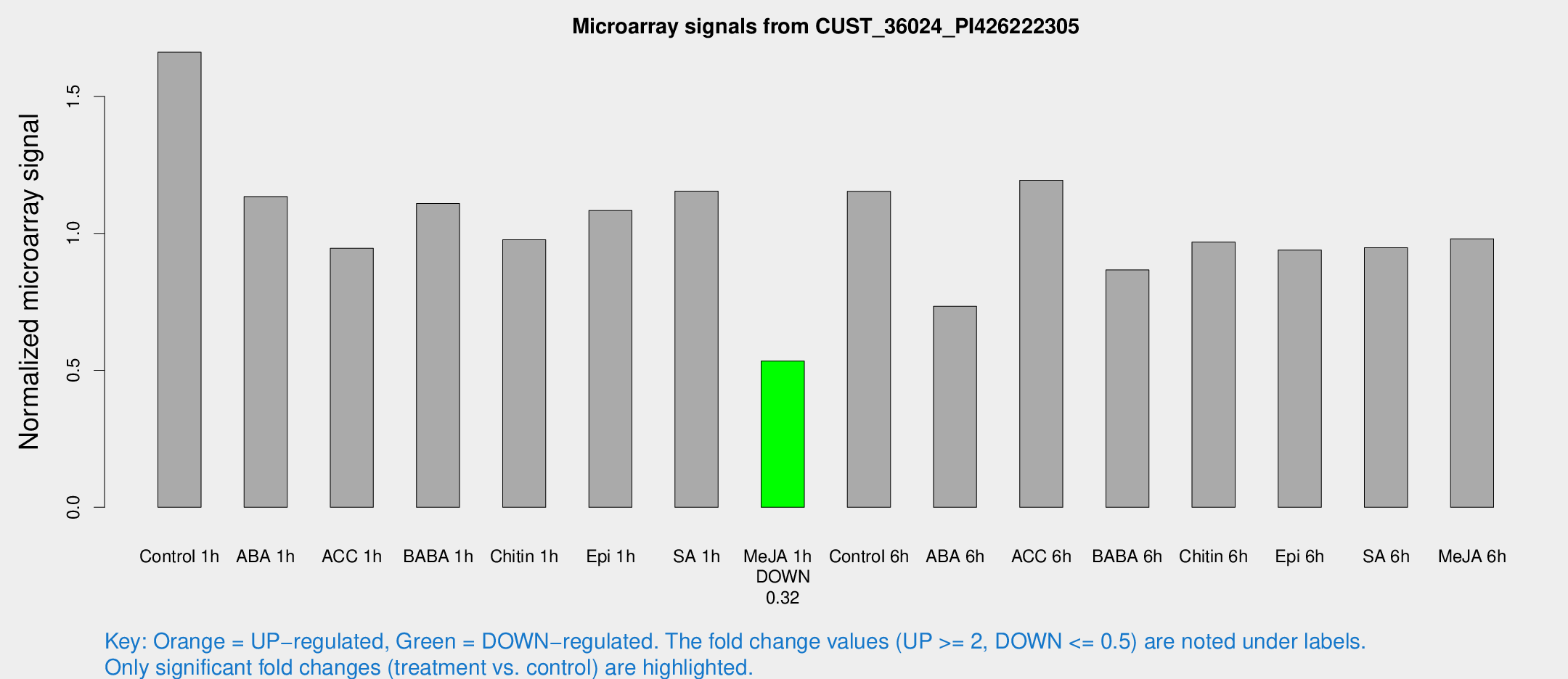
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1829.98 | 173.979 | 1.66143 | 0.0982897 |
| ABA 1h | 1104.39 | 96.7581 | 1.1342 | 0.0655612 |
| ACC 1h | 1108.47 | 213.755 | 0.945407 | 0.136798 |
| BABA 1h | 1225.83 | 245.437 | 1.10917 | 0.139558 |
| Chitin 1h | 963.537 | 64.4604 | 0.976805 | 0.0564881 |
| Epi 1h | 1032.61 | 90.9921 | 1.08358 | 0.0893469 |
| SA 1h | 1309.28 | 142.16 | 1.15398 | 0.0666834 |
| Me-JA 1h | 483.447 | 58.6701 | 0.534001 | 0.0310463 |
| Control 6h | 1328.09 | 281.308 | 1.15364 | 0.177643 |
| ABA 6h | 884.706 | 172.621 | 0.733646 | 0.103502 |
| ACC 6h | 1504.83 | 117.93 | 1.19397 | 0.0806836 |
| BABA 6h | 1066.45 | 84.764 | 0.866658 | 0.0501295 |
| Chitin 6h | 1145.89 | 147.678 | 0.968009 | 0.143666 |
| Epi 6h | 1160.46 | 68.122 | 0.939143 | 0.0543112 |
| SA 6h | 1099.95 | 273.979 | 0.947786 | 0.160655 |
| Me-JA 6h | 1106.87 | 223.759 | 0.980456 | 0.124123 |
Source Transcript PGSC0003DMT400080928 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G31500.1 | +2 | 2e-149 | 443 | 236/495 (48%) | cytochrome P450, family 83, subfamily B, polypeptide 1 | chr4:15273677-15275271 REVERSE LENGTH=499 |
