Probe CUST_36021_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36021_PI426222305 | JHI_St_60k_v1 | DMT400080927 | GGGCGAGTTCTGTAAGGTTAATTAAAGTCGTTACATAATGCCACATCATGTATTATTTAC |
All Microarray Probes Designed to Gene DMG402031520
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36024_PI426222305 | JHI_St_60k_v1 | DMT400080928 | GGGCGAGTTCTGTAAGGTTAATTAAAGTCGTTACATAATGCCACATCATGTATTATTTAC |
| CUST_36021_PI426222305 | JHI_St_60k_v1 | DMT400080927 | GGGCGAGTTCTGTAAGGTTAATTAAAGTCGTTACATAATGCCACATCATGTATTATTTAC |
Microarray Signals from CUST_36021_PI426222305
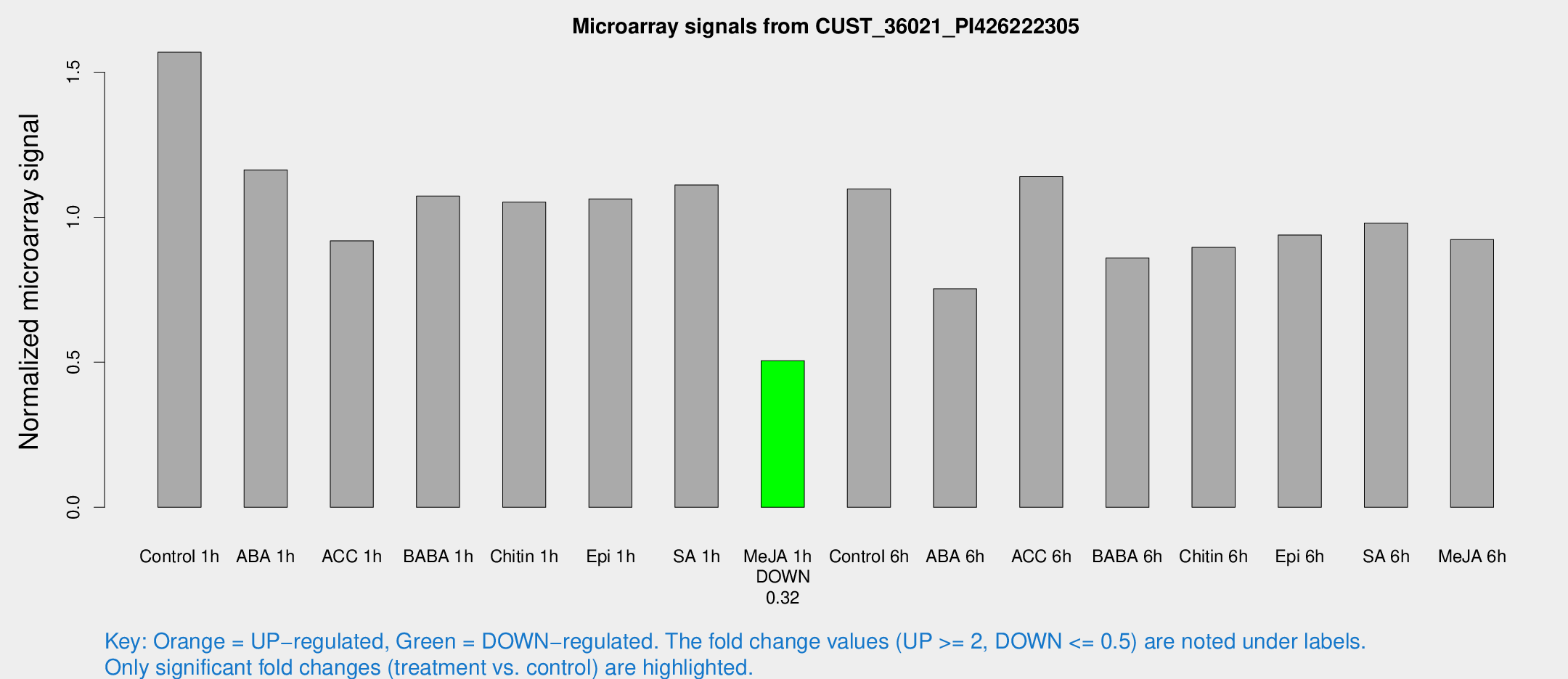
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1713.95 | 232.968 | 1.56858 | 0.126298 |
| ABA 1h | 1109.69 | 70.3303 | 1.16319 | 0.0672604 |
| ACC 1h | 1053.08 | 192.235 | 0.918646 | 0.119816 |
| BABA 1h | 1152.23 | 203.719 | 1.0727 | 0.100262 |
| Chitin 1h | 1019.78 | 60.7024 | 1.05216 | 0.0608669 |
| Epi 1h | 1001.11 | 112.248 | 1.06291 | 0.118104 |
| SA 1h | 1239.86 | 134.126 | 1.11083 | 0.0642191 |
| Me-JA 1h | 446.041 | 34.2666 | 0.505584 | 0.0294873 |
| Control 6h | 1232.05 | 248.788 | 1.09708 | 0.157822 |
| ABA 6h | 886.887 | 158.79 | 0.753176 | 0.090179 |
| ACC 6h | 1420.24 | 146.134 | 1.13956 | 0.117233 |
| BABA 6h | 1049.67 | 125.654 | 0.859379 | 0.0796214 |
| Chitin 6h | 1040.48 | 124.093 | 0.895901 | 0.118173 |
| Epi 6h | 1146.6 | 103.291 | 0.938649 | 0.054317 |
| SA 6h | 1109.93 | 260.943 | 0.980035 | 0.149057 |
| Me-JA 6h | 1032.2 | 225.815 | 0.922947 | 0.124269 |
Source Transcript PGSC0003DMT400080927 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G31500.1 | +1 | 2e-66 | 229 | 115/201 (57%) | cytochrome P450, family 83, subfamily B, polypeptide 1 | chr4:15273677-15275271 REVERSE LENGTH=499 |
