Probe CUST_35995_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35995_PI426222305 | JHI_St_60k_v1 | DMT400080920 | TGGTTAAGTTCTTCCAAATTTATGTAGGATGTGTTCGTTGCTGGATCGGACACTAGCGCG |
All Microarray Probes Designed to Gene DMG400031518
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35936_PI426222305 | JHI_St_60k_v1 | DMT400080919 | TTTGGATGACATCAAAGGAATACTTATGGATGTGTTCGTTGCTGGATCGGACACTAGCGC |
| CUST_35995_PI426222305 | JHI_St_60k_v1 | DMT400080920 | TGGTTAAGTTCTTCCAAATTTATGTAGGATGTGTTCGTTGCTGGATCGGACACTAGCGCG |
Microarray Signals from CUST_35995_PI426222305
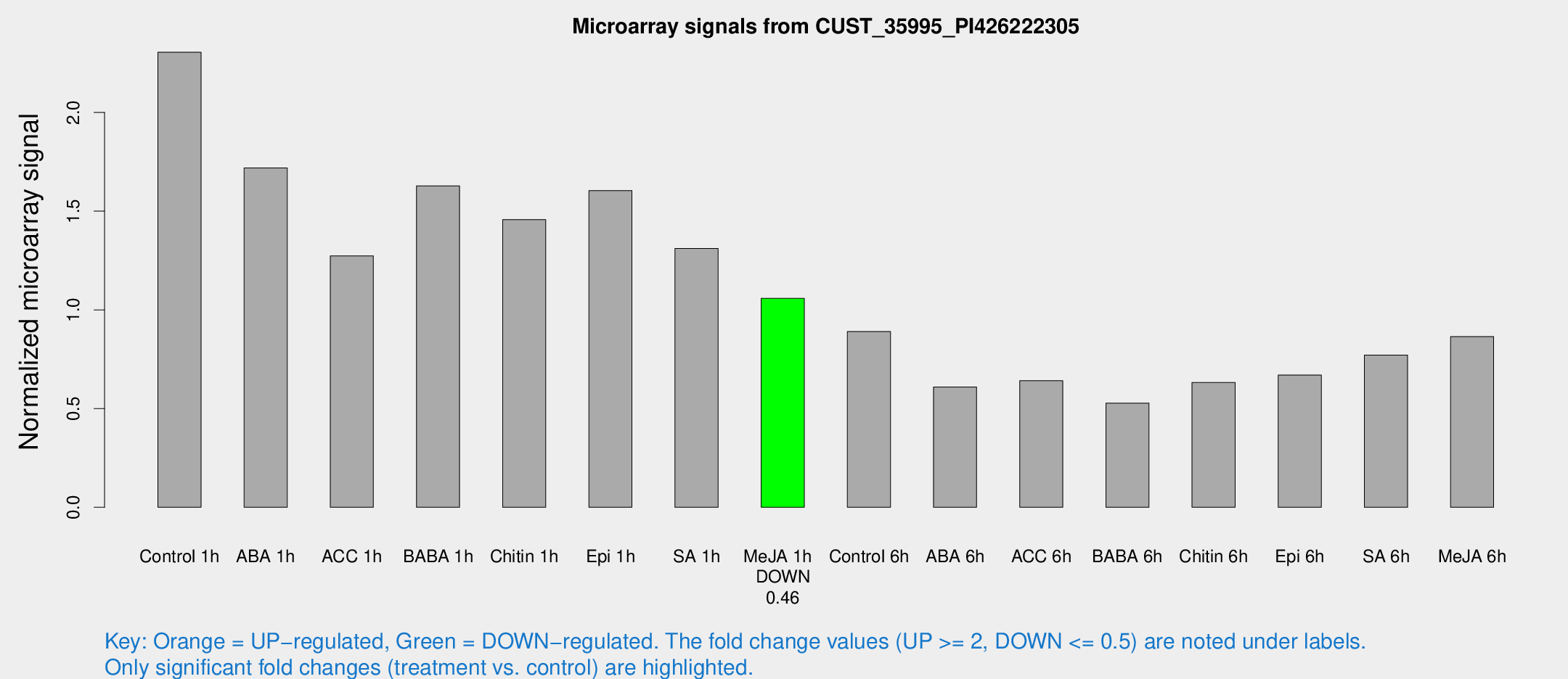
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1148.93 | 66.6374 | 2.30471 | 0.133254 |
| ABA 1h | 768.334 | 94.6814 | 1.71873 | 0.131481 |
| ACC 1h | 693.244 | 158.015 | 1.27311 | 0.249768 |
| BABA 1h | 848.114 | 221.009 | 1.62795 | 0.338383 |
| Chitin 1h | 661.121 | 82.5774 | 1.4574 | 0.084544 |
| Epi 1h | 692.381 | 40.9457 | 1.60423 | 0.116325 |
| SA 1h | 694.14 | 123.272 | 1.31058 | 0.189137 |
| Me-JA 1h | 449.873 | 98.777 | 1.05815 | 0.137766 |
| Control 6h | 478.282 | 122.212 | 0.890266 | 0.187447 |
| ABA 6h | 345.084 | 81.0211 | 0.609456 | 0.151132 |
| ACC 6h | 374.675 | 55.1296 | 0.641341 | 0.073634 |
| BABA 6h | 310.197 | 76.0879 | 0.527795 | 0.119578 |
| Chitin 6h | 347.861 | 65.1244 | 0.632213 | 0.139843 |
| Epi 6h | 398.809 | 103.074 | 0.669718 | 0.163973 |
| SA 6h | 385.539 | 52.777 | 0.770682 | 0.202305 |
| Me-JA 6h | 438.059 | 68.499 | 0.864541 | 0.0646872 |
Source Transcript PGSC0003DMT400080920 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G26330.1 | +2 | 5e-74 | 237 | 110/201 (55%) | cytochrome P450, family 71, subfamily B, polypeptide 37 | chr3:9646873-9648536 REVERSE LENGTH=500 |
