Probe CUST_35936_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35936_PI426222305 | JHI_St_60k_v1 | DMT400080919 | TTTGGATGACATCAAAGGAATACTTATGGATGTGTTCGTTGCTGGATCGGACACTAGCGC |
All Microarray Probes Designed to Gene DMG400031518
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35936_PI426222305 | JHI_St_60k_v1 | DMT400080919 | TTTGGATGACATCAAAGGAATACTTATGGATGTGTTCGTTGCTGGATCGGACACTAGCGC |
| CUST_35995_PI426222305 | JHI_St_60k_v1 | DMT400080920 | TGGTTAAGTTCTTCCAAATTTATGTAGGATGTGTTCGTTGCTGGATCGGACACTAGCGCG |
Microarray Signals from CUST_35936_PI426222305
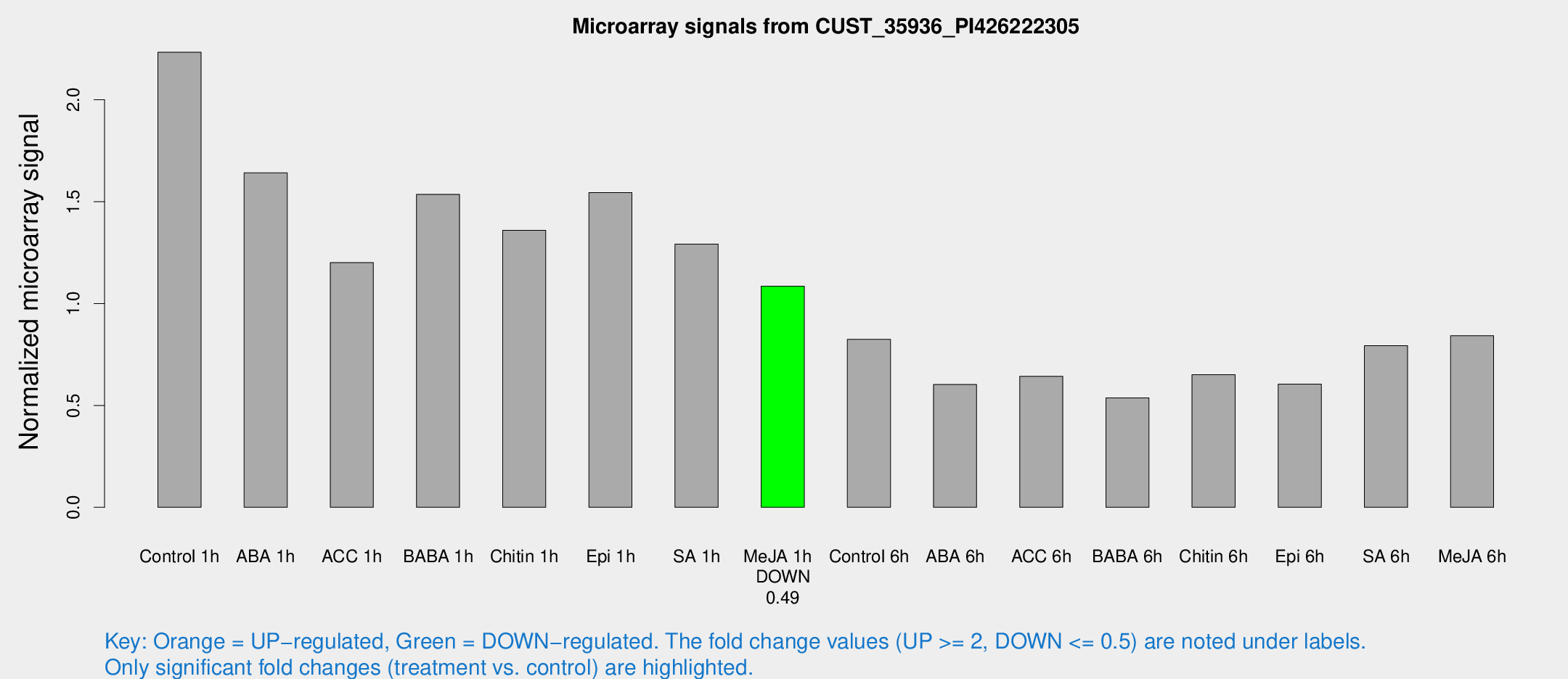
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1702.06 | 115.659 | 2.23358 | 0.129035 |
| ABA 1h | 1122.64 | 148.57 | 1.64141 | 0.131474 |
| ACC 1h | 981.577 | 195.198 | 1.20095 | 0.182964 |
| BABA 1h | 1235.67 | 343.984 | 1.53609 | 0.355836 |
| Chitin 1h | 933.006 | 73.5454 | 1.35973 | 0.0786634 |
| Epi 1h | 1020.7 | 85.0068 | 1.54431 | 0.153351 |
| SA 1h | 1043.35 | 183.712 | 1.29226 | 0.178517 |
| Me-JA 1h | 692.034 | 121.019 | 1.08525 | 0.0929601 |
| Control 6h | 684.262 | 198.78 | 0.824201 | 0.199578 |
| ABA 6h | 546.91 | 158.952 | 0.602507 | 0.215508 |
| ACC 6h | 573.236 | 82.8517 | 0.643352 | 0.0896736 |
| BABA 6h | 490.791 | 138.659 | 0.536968 | 0.141488 |
| Chitin 6h | 530.725 | 49.6073 | 0.651142 | 0.0687696 |
| Epi 6h | 543.919 | 127.626 | 0.604612 | 0.132744 |
| SA 6h | 598.966 | 49.0347 | 0.79314 | 0.162079 |
| Me-JA 6h | 658.653 | 123.282 | 0.842339 | 0.0873661 |
Source Transcript PGSC0003DMT400080919 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G31500.1 | +1 | 1e-128 | 389 | 198/407 (49%) | cytochrome P450, family 83, subfamily B, polypeptide 1 | chr4:15273677-15275271 REVERSE LENGTH=499 |
