Probe CUST_35583_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35583_PI426222305 | JHI_St_60k_v1 | DMT400004649 | ACATCTCCTCATGCATCATTAAACAAGTTTCCTCTGTTCCTTCTCAGCTAACTAATGTAT |
All Microarray Probes Designed to Gene DMG400001844
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35556_PI426222305 | JHI_St_60k_v1 | DMT400004650 | CACGTGCTTATAAAGTTATAATAGGTGTATTACACACTATGCCATGTCAACAATGTGTCT |
| CUST_35583_PI426222305 | JHI_St_60k_v1 | DMT400004649 | ACATCTCCTCATGCATCATTAAACAAGTTTCCTCTGTTCCTTCTCAGCTAACTAATGTAT |
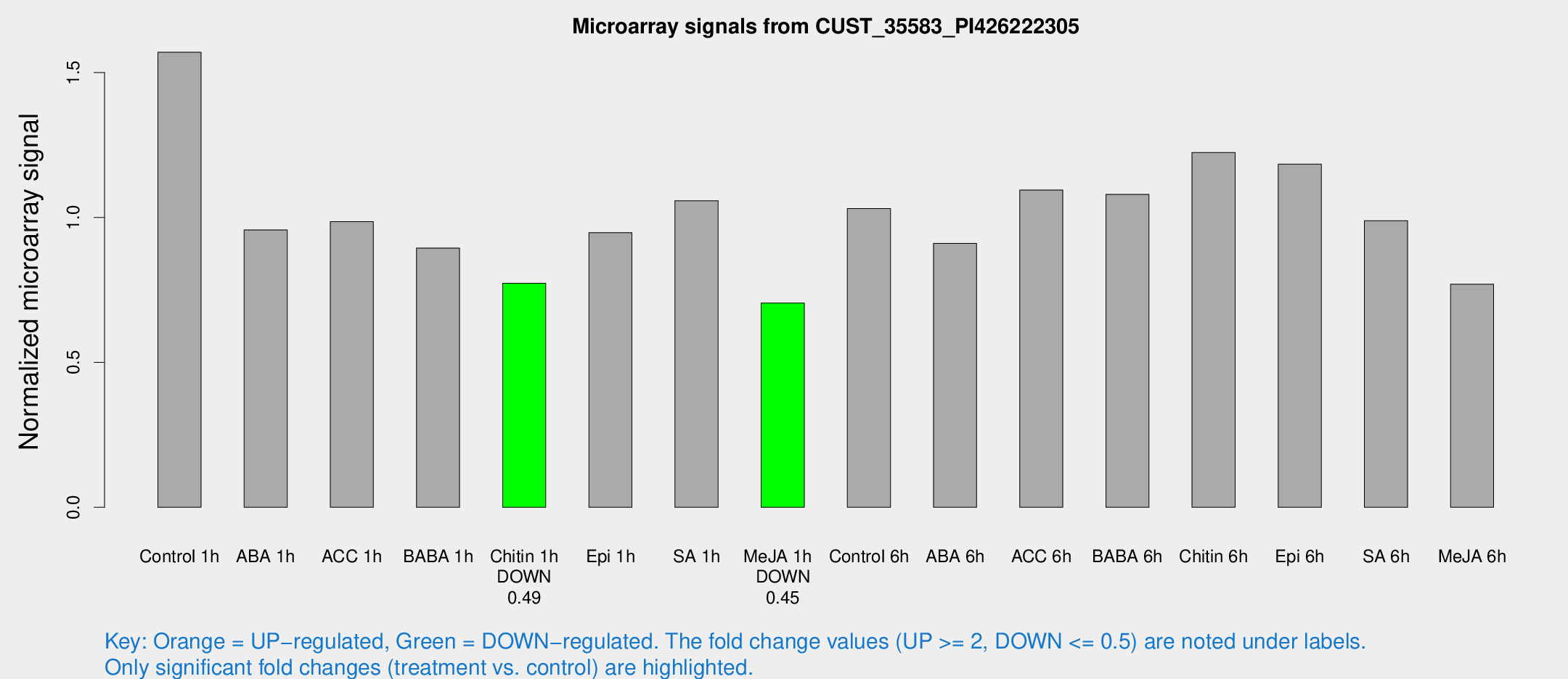
Microarray Signals from CUST_35583_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 606.872 | 44.6108 | 1.5703 | 0.0910521 |
| ABA 1h | 326.905 | 20.846 | 0.957276 | 0.0560265 |
| ACC 1h | 428.054 | 114.166 | 0.986021 | 0.257417 |
| BABA 1h | 337.369 | 44.9707 | 0.894499 | 0.0646277 |
| Chitin 1h | 267.314 | 15.7725 | 0.773186 | 0.0595845 |
| Epi 1h | 320.999 | 41.3313 | 0.947753 | 0.122466 |
| SA 1h | 422.847 | 46.0211 | 1.05767 | 0.16785 |
| Me-JA 1h | 221.639 | 13.2391 | 0.704644 | 0.0639469 |
| Control 6h | 410.084 | 73.6281 | 1.0309 | 0.104494 |
| ABA 6h | 376.665 | 39.8164 | 0.91073 | 0.124152 |
| ACC 6h | 491.039 | 63.9085 | 1.09499 | 0.0638378 |
| BABA 6h | 474.618 | 68.5995 | 1.07978 | 0.133618 |
| Chitin 6h | 503.2 | 32.1478 | 1.22443 | 0.119388 |
| Epi 6h | 513.492 | 29.9007 | 1.1843 | 0.0732126 |
| SA 6h | 381.921 | 48.6488 | 0.989096 | 0.249955 |
| Me-JA 6h | 311.457 | 75.7567 | 0.770189 | 0.121696 |
Source Transcript PGSC0003DMT400004649 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G38610.1 | +1 | 2e-15 | 70 | 43/126 (34%) | Plant invertase/pectin methylesterase inhibitor superfamily protein | chr5:15460845-15461444 REVERSE LENGTH=199 |
