Probe CUST_35556_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35556_PI426222305 | JHI_St_60k_v1 | DMT400004650 | CACGTGCTTATAAAGTTATAATAGGTGTATTACACACTATGCCATGTCAACAATGTGTCT |
All Microarray Probes Designed to Gene DMG400001844
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35556_PI426222305 | JHI_St_60k_v1 | DMT400004650 | CACGTGCTTATAAAGTTATAATAGGTGTATTACACACTATGCCATGTCAACAATGTGTCT |
| CUST_35583_PI426222305 | JHI_St_60k_v1 | DMT400004649 | ACATCTCCTCATGCATCATTAAACAAGTTTCCTCTGTTCCTTCTCAGCTAACTAATGTAT |
Microarray Signals from CUST_35556_PI426222305
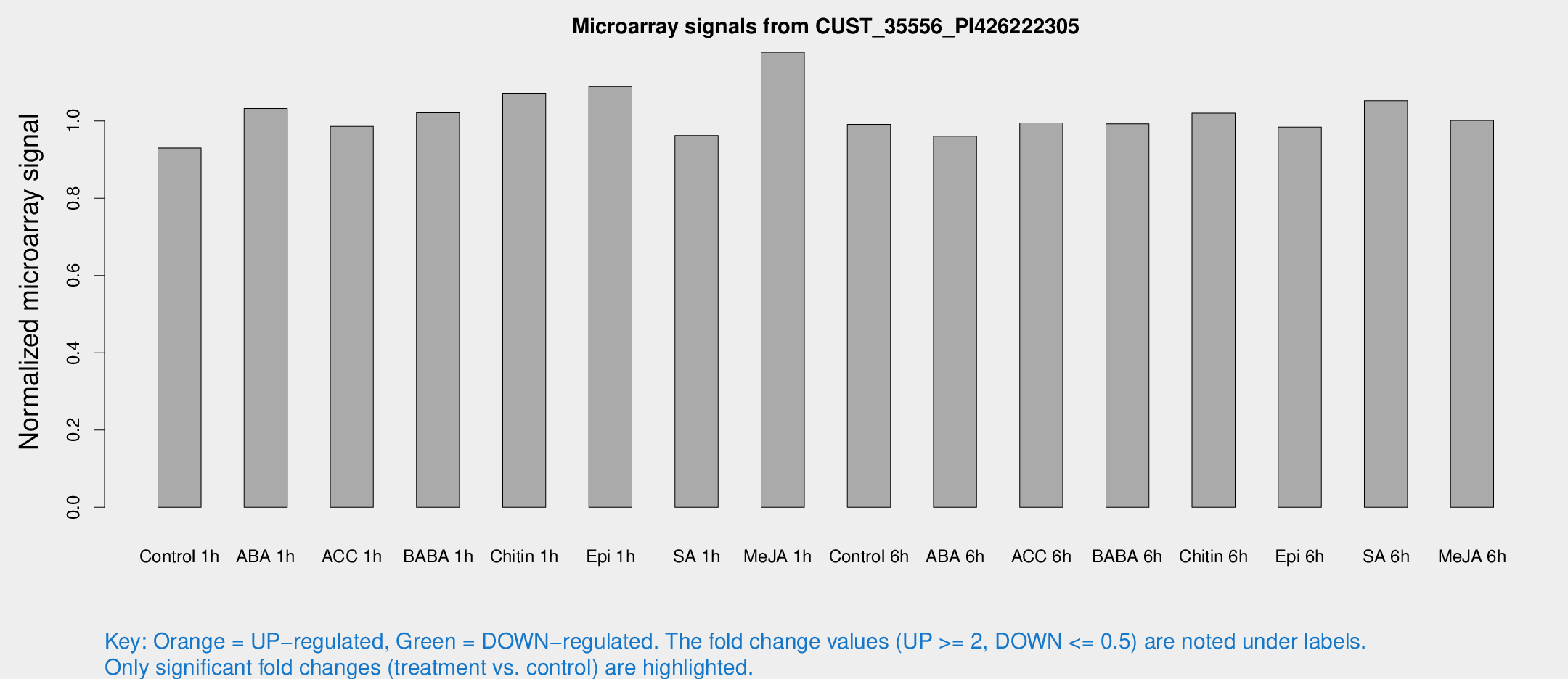
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.17241 | 2.99738 | 0.9301 | 0.534489 |
| ABA 1h | 5.07424 | 2.94047 | 1.03221 | 0.592851 |
| ACC 1h | 5.67121 | 3.28823 | 0.985633 | 0.571169 |
| BABA 1h | 5.52064 | 3.20217 | 1.02112 | 0.591287 |
| Chitin 1h | 5.29116 | 3.07963 | 1.07182 | 0.60578 |
| Epi 1h | 5.20555 | 3.02066 | 1.08921 | 0.620799 |
| SA 1h | 5.54747 | 3.04645 | 0.962142 | 0.530745 |
| Me-JA 1h | 5.402 | 3.14102 | 1.17769 | 0.682484 |
| Control 6h | 5.56442 | 3.22369 | 0.991038 | 0.573905 |
| ABA 6h | 5.72216 | 3.31816 | 0.960425 | 0.556498 |
| ACC 6h | 6.52641 | 3.86446 | 0.99461 | 0.575949 |
| BABA 6h | 6.23812 | 3.62273 | 0.992414 | 0.574688 |
| Chitin 6h | 6.0804 | 3.52142 | 1.01985 | 0.590516 |
| Epi 6h | 6.26689 | 3.66502 | 0.98368 | 0.570095 |
| SA 6h | 5.83118 | 3.37979 | 1.0523 | 0.609658 |
| Me-JA 6h | 5.58736 | 3.23935 | 1.00115 | 0.579961 |
Source Transcript PGSC0003DMT400004650 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G38610.1 | +2 | 9e-26 | 108 | 65/175 (37%) | Plant invertase/pectin methylesterase inhibitor superfamily protein | chr5:15460845-15461444 REVERSE LENGTH=199 |
