Probe CUST_35508_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35508_PI426222305 | JHI_St_60k_v1 | DMT400005816 | TGATCCAAATTTGAACTGTCAAGCCAGTTAGCATCACAATCTGTATTGTCTTGTGTATAT |
All Microarray Probes Designed to Gene DMG403002270
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35482_PI426222305 | JHI_St_60k_v1 | DMT400005815 | AACTGGCATTGATACTGAATTTGTGGAGGTTATGGCTTTCAGGCATGTTCATGATGTGGC |
| CUST_35508_PI426222305 | JHI_St_60k_v1 | DMT400005816 | TGATCCAAATTTGAACTGTCAAGCCAGTTAGCATCACAATCTGTATTGTCTTGTGTATAT |
Microarray Signals from CUST_35508_PI426222305
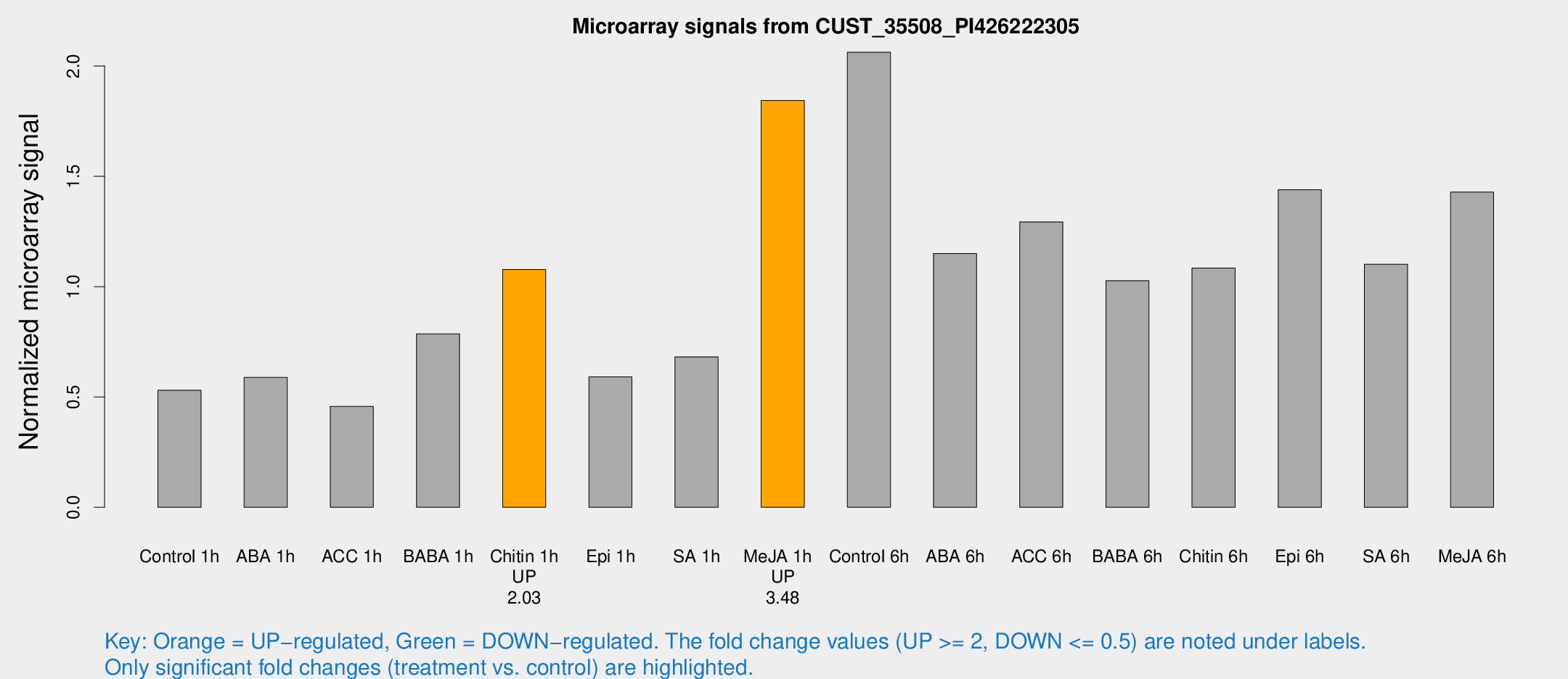
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 4763.7 | 301.554 | 0.530464 | 0.050853 |
| ABA 1h | 4920.47 | 1195.85 | 0.588476 | 0.112002 |
| ACC 1h | 4279.32 | 595.618 | 0.457234 | 0.0733126 |
| BABA 1h | 7138.88 | 1533.28 | 0.785685 | 0.109482 |
| Chitin 1h | 8673.47 | 501.318 | 1.07746 | 0.08614 |
| Epi 1h | 4657.58 | 593.475 | 0.5916 | 0.0654784 |
| SA 1h | 6415.04 | 1023.42 | 0.681945 | 0.106638 |
| Me-JA 1h | 13754.7 | 1931.29 | 1.84395 | 0.12894 |
| Control 6h | 19382.1 | 4173.66 | 2.0624 | 0.279657 |
| ABA 6h | 11179.3 | 1636.09 | 1.15042 | 0.107807 |
| ACC 6h | 13474.8 | 1677.68 | 1.29349 | 0.074681 |
| BABA 6h | 10392.6 | 1109.74 | 1.02705 | 0.106435 |
| Chitin 6h | 10359.7 | 617.494 | 1.08423 | 0.100167 |
| Epi 6h | 15262.9 | 3366.74 | 1.43977 | 0.404085 |
| SA 6h | 10255.3 | 2142.69 | 1.10176 | 0.135553 |
| Me-JA 6h | 13187.1 | 2395.96 | 1.42854 | 0.155825 |
Source Transcript PGSC0003DMT400005816 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G47240.1 | +1 | 1e-14 | 71 | 40/61 (66%) | nudix hydrolase homolog 8 | chr5:19183806-19185467 FORWARD LENGTH=369 |
