Probe CUST_35482_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35482_PI426222305 | JHI_St_60k_v1 | DMT400005815 | AACTGGCATTGATACTGAATTTGTGGAGGTTATGGCTTTCAGGCATGTTCATGATGTGGC |
All Microarray Probes Designed to Gene DMG403002270
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35482_PI426222305 | JHI_St_60k_v1 | DMT400005815 | AACTGGCATTGATACTGAATTTGTGGAGGTTATGGCTTTCAGGCATGTTCATGATGTGGC |
| CUST_35508_PI426222305 | JHI_St_60k_v1 | DMT400005816 | TGATCCAAATTTGAACTGTCAAGCCAGTTAGCATCACAATCTGTATTGTCTTGTGTATAT |
Microarray Signals from CUST_35482_PI426222305
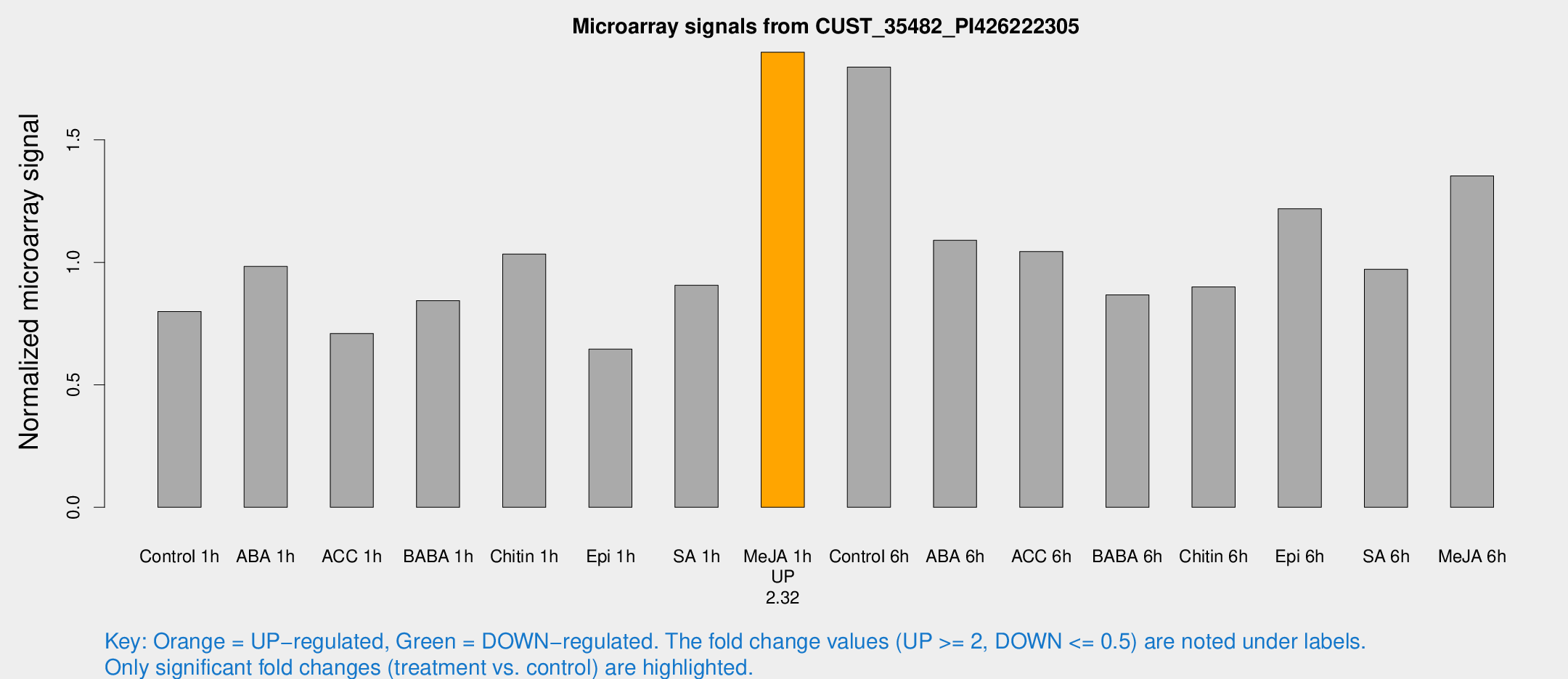
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9466.02 | 587.974 | 0.799477 | 0.0461588 |
| ABA 1h | 10740.2 | 2407.29 | 0.983231 | 0.148325 |
| ACC 1h | 8829.07 | 1405.6 | 0.70919 | 0.0734709 |
| BABA 1h | 10161.2 | 2236.09 | 0.84383 | 0.125628 |
| Chitin 1h | 11041.4 | 883.186 | 1.03389 | 0.0596926 |
| Epi 1h | 6745.62 | 978.777 | 0.645636 | 0.0882067 |
| SA 1h | 11586.1 | 2724.04 | 0.906573 | 0.19938 |
| Me-JA 1h | 18299 | 2670.82 | 1.8579 | 0.124384 |
| Control 6h | 22267.8 | 4526.99 | 1.79723 | 0.232716 |
| ABA 6h | 13919 | 1858.39 | 1.09013 | 0.0857778 |
| ACC 6h | 14411.1 | 1995.31 | 1.04387 | 0.0696282 |
| BABA 6h | 11610.6 | 1426.3 | 0.866882 | 0.0986275 |
| Chitin 6h | 11362.6 | 840.234 | 0.89969 | 0.0975884 |
| Epi 6h | 17054.9 | 3835.69 | 1.21863 | 0.337903 |
| SA 6h | 12439.4 | 3273.24 | 0.971867 | 0.206019 |
| Me-JA 6h | 16483.7 | 3064.36 | 1.35308 | 0.146795 |
Source Transcript PGSC0003DMT400005815 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G47240.1 | +3 | 1e-42 | 146 | 75/111 (68%) | nudix hydrolase homolog 8 | chr5:19183806-19185467 FORWARD LENGTH=369 |
