Probe CUST_34699_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34699_PI426222305 | JHI_St_60k_v1 | DMT400001780 | CCTTCTCACCCTTATGATAGATCCCAAGCCAGATTTTGGGCTGACTATATAGATAAAAAG |
All Microarray Probes Designed to Gene DMG400000667
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34699_PI426222305 | JHI_St_60k_v1 | DMT400001780 | CCTTCTCACCCTTATGATAGATCCCAAGCCAGATTTTGGGCTGACTATATAGATAAAAAG |
| CUST_34683_PI426222305 | JHI_St_60k_v1 | DMT400001779 | TGGAACTCAAGAACAAATTTGGTCTACCCTAGATGGAAGGCACCAATAATGTGGTTCTAC |
Microarray Signals from CUST_34699_PI426222305
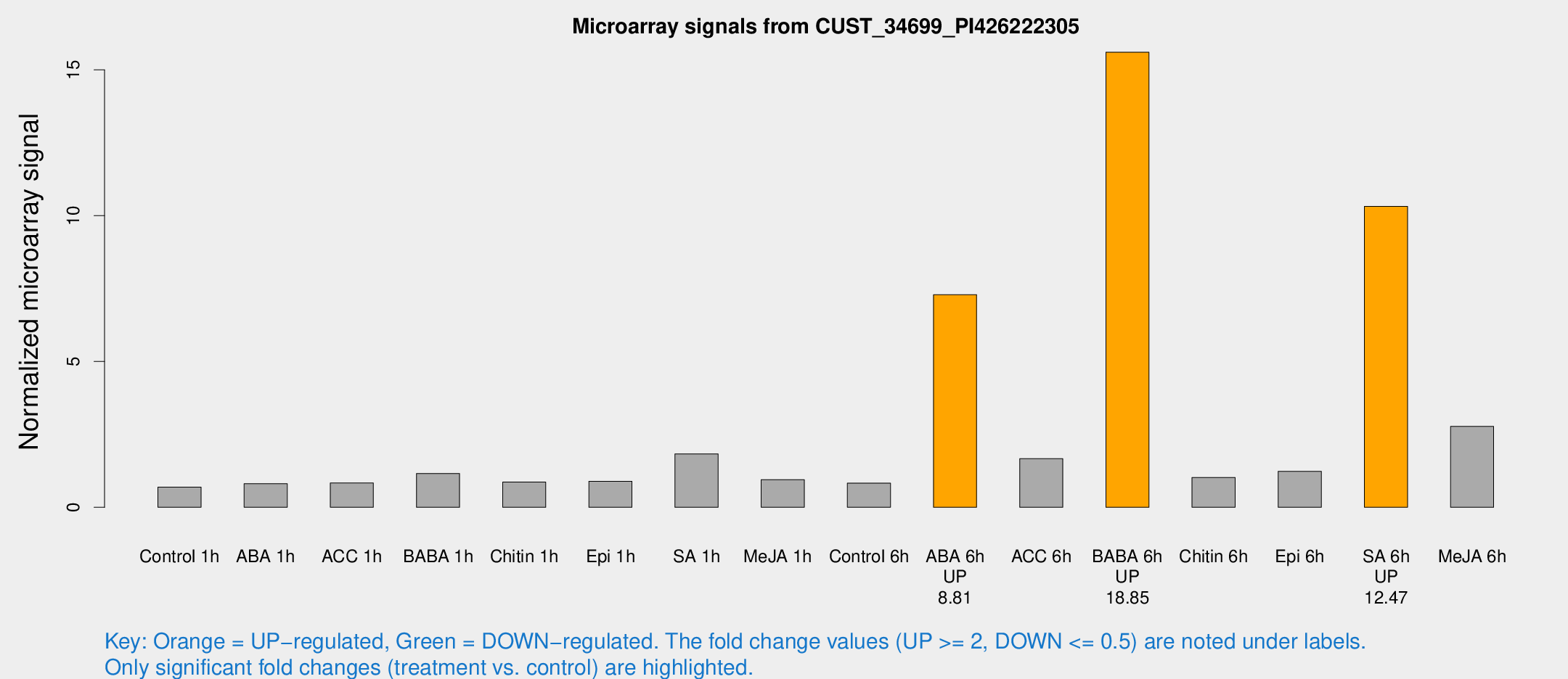
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.15122 | 3.57424 | 0.694164 | 0.402025 |
| ABA 1h | 6.34591 | 3.43404 | 0.808368 | 0.440969 |
| ACC 1h | 7.77736 | 4.64633 | 0.832799 | 0.482646 |
| BABA 1h | 11.3408 | 4.37978 | 1.16098 | 0.499341 |
| Chitin 1h | 7.22233 | 3.58207 | 0.867034 | 0.451069 |
| Epi 1h | 6.87897 | 3.45136 | 0.893424 | 0.458246 |
| SA 1h | 20.6924 | 7.57518 | 1.83065 | 1.03794 |
| Me-JA 1h | 7.06452 | 3.7054 | 0.947888 | 0.509008 |
| Control 6h | 7.36375 | 3.76638 | 0.827734 | 0.425535 |
| ABA 6h | 85.4341 | 37.0043 | 7.29166 | 3.69353 |
| ACC 6h | 21.229 | 10.5246 | 1.66573 | 1.23359 |
| BABA 6h | 158.527 | 26.9956 | 15.6057 | 3.18507 |
| Chitin 6h | 10.6533 | 4.1611 | 1.02308 | 0.486143 |
| Epi 6h | 12.4769 | 4.41625 | 1.23235 | 0.471711 |
| SA 6h | 93.1406 | 17.5377 | 10.3206 | 1.71966 |
| Me-JA 6h | 35.272 | 17.8869 | 2.77451 | 3.02512 |
Source Transcript PGSC0003DMT400001780 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G17180.1 | +2 | 2e-68 | 204 | 95/158 (60%) | glutathione S-transferase TAU 25 | chr1:5872208-5872958 FORWARD LENGTH=221 |
