Probe CUST_34683_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34683_PI426222305 | JHI_St_60k_v1 | DMT400001779 | TGGAACTCAAGAACAAATTTGGTCTACCCTAGATGGAAGGCACCAATAATGTGGTTCTAC |
All Microarray Probes Designed to Gene DMG400000667
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34699_PI426222305 | JHI_St_60k_v1 | DMT400001780 | CCTTCTCACCCTTATGATAGATCCCAAGCCAGATTTTGGGCTGACTATATAGATAAAAAG |
| CUST_34683_PI426222305 | JHI_St_60k_v1 | DMT400001779 | TGGAACTCAAGAACAAATTTGGTCTACCCTAGATGGAAGGCACCAATAATGTGGTTCTAC |
Microarray Signals from CUST_34683_PI426222305
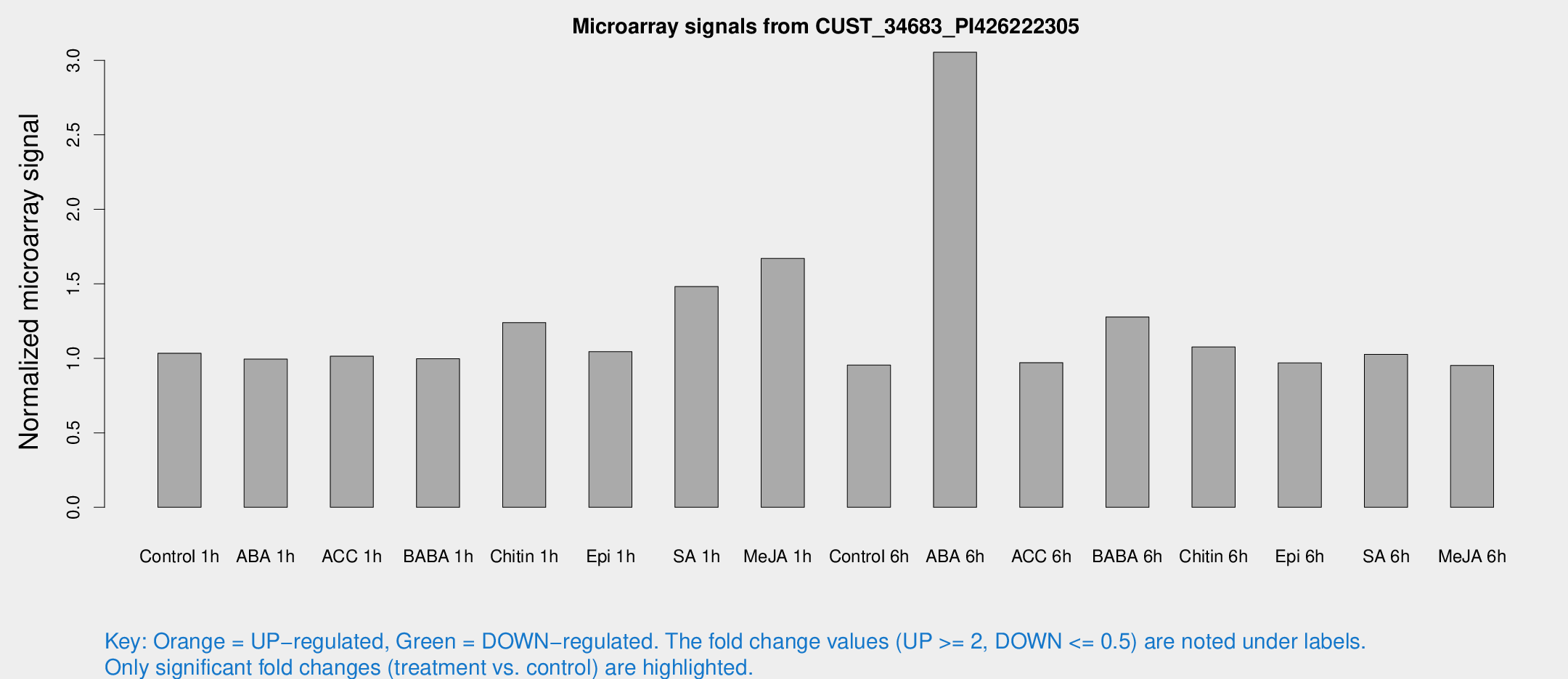
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.27194 | 3.20035 | 1.03386 | 0.542919 |
| ABA 1h | 5.24941 | 3.04522 | 0.994613 | 0.576406 |
| ACC 1h | 6.25274 | 3.64977 | 1.01402 | 0.587901 |
| BABA 1h | 5.73811 | 3.33291 | 0.997134 | 0.578662 |
| Chitin 1h | 6.81784 | 3.18994 | 1.23976 | 0.617327 |
| Epi 1h | 5.392 | 3.13137 | 1.04457 | 0.604358 |
| SA 1h | 9.70509 | 3.19286 | 1.48121 | 0.569935 |
| Me-JA 1h | 8.62821 | 3.28212 | 1.67035 | 0.719889 |
| Control 6h | 5.71046 | 3.30769 | 0.955074 | 0.553079 |
| ABA 6h | 31.1242 | 20.7445 | 3.05417 | 2.64797 |
| ACC 6h | 6.80106 | 4.0404 | 0.971001 | 0.563127 |
| BABA 6h | 8.8738 | 3.66802 | 1.27741 | 0.579807 |
| Chitin 6h | 6.87465 | 3.68233 | 1.07646 | 0.582796 |
| Epi 6h | 6.57026 | 3.83993 | 0.96917 | 0.561693 |
| SA 6h | 6.0604 | 3.45846 | 1.02666 | 0.586052 |
| Me-JA 6h | 5.66221 | 3.28318 | 0.952219 | 0.551502 |
Source Transcript PGSC0003DMT400001779 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G78380.1 | +1 | 1e-57 | 183 | 99/215 (46%) | glutathione S-transferase TAU 19 | chr1:29486659-29487819 REVERSE LENGTH=219 |
