Probe CUST_33237_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33237_PI426222305 | JHI_St_60k_v1 | DMT400067599 | CTGAAGGAAATTTACATGGCATTTTGGTTGGGTAAGTATCAATTTGATTTTGAGCCATCA |
All Microarray Probes Designed to Gene DMG400026280
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33237_PI426222305 | JHI_St_60k_v1 | DMT400067599 | CTGAAGGAAATTTACATGGCATTTTGGTTGGGTAAGTATCAATTTGATTTTGAGCCATCA |
| CUST_33144_PI426222305 | JHI_St_60k_v1 | DMT400067600 | GGTGATGAAGGGCTTCTACCATAGAGTTTGTCTACATATTACAACTCTTTATGACTTTAT |
Microarray Signals from CUST_33237_PI426222305
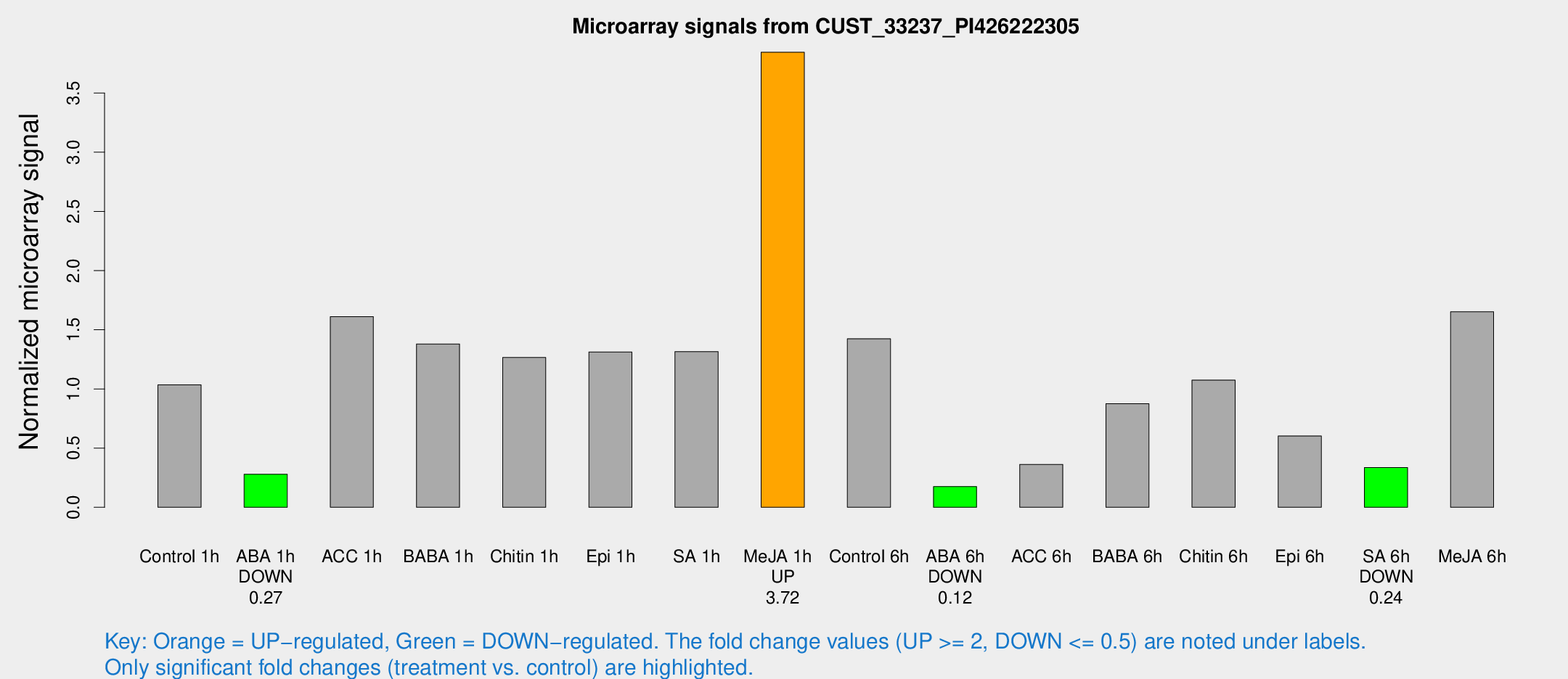
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 217.572 | 45.7107 | 1.03485 | 0.143222 |
| ABA 1h | 57.6123 | 21.3507 | 0.278715 | 0.0967544 |
| ACC 1h | 372.849 | 116.488 | 1.61104 | 0.475453 |
| BABA 1h | 269.671 | 19.815 | 1.37994 | 0.0918347 |
| Chitin 1h | 239.041 | 47.0295 | 1.26559 | 0.223668 |
| Epi 1h | 234.537 | 36.2537 | 1.31209 | 0.172056 |
| SA 1h | 300.898 | 98.9105 | 1.3158 | 0.421743 |
| Me-JA 1h | 656.506 | 130.508 | 3.84467 | 0.427255 |
| Control 6h | 359.976 | 175.92 | 1.4243 | 0.628648 |
| ABA 6h | 40.1232 | 10.5279 | 0.17545 | 0.0383161 |
| ACC 6h | 85.0482 | 10.3467 | 0.362531 | 0.025597 |
| BABA 6h | 203.502 | 32.4469 | 0.875373 | 0.123968 |
| Chitin 6h | 239.068 | 46.9691 | 1.07483 | 0.204035 |
| Epi 6h | 163.74 | 55.5839 | 0.602955 | 0.402954 |
| SA 6h | 75.8363 | 27.9396 | 0.33574 | 0.101543 |
| Me-JA 6h | 348.062 | 69.7626 | 1.65205 | 0.247223 |
Source Transcript PGSC0003DMT400067599 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G43710.1 | +1 | 2e-36 | 133 | 75/166 (45%) | Pyridoxal phosphate (PLP)-dependent transferases superfamily protein | chr1:16486534-16488298 REVERSE LENGTH=482 |
