Probe CUST_33144_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33144_PI426222305 | JHI_St_60k_v1 | DMT400067600 | GGTGATGAAGGGCTTCTACCATAGAGTTTGTCTACATATTACAACTCTTTATGACTTTAT |
All Microarray Probes Designed to Gene DMG400026280
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33237_PI426222305 | JHI_St_60k_v1 | DMT400067599 | CTGAAGGAAATTTACATGGCATTTTGGTTGGGTAAGTATCAATTTGATTTTGAGCCATCA |
| CUST_33144_PI426222305 | JHI_St_60k_v1 | DMT400067600 | GGTGATGAAGGGCTTCTACCATAGAGTTTGTCTACATATTACAACTCTTTATGACTTTAT |
Microarray Signals from CUST_33144_PI426222305
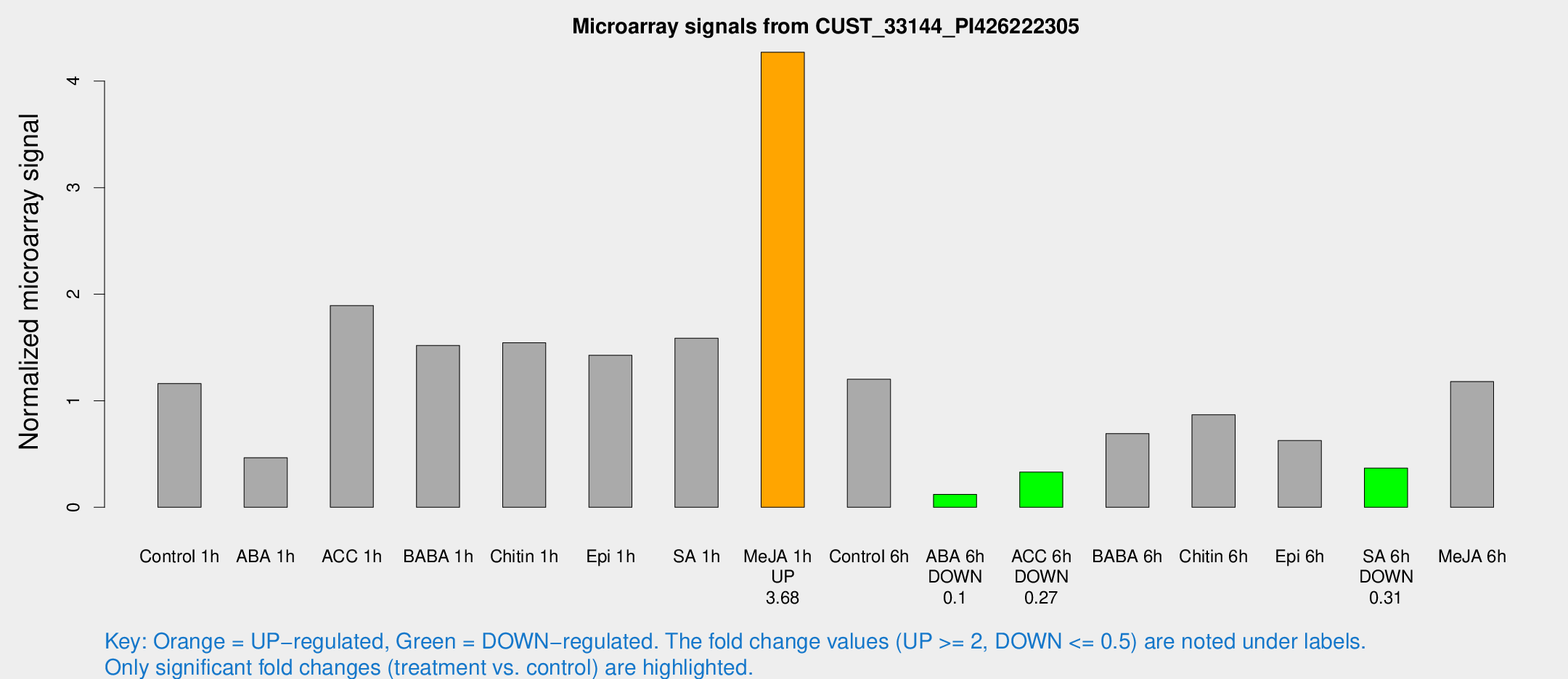
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11455 | 691.328 | 1.16209 | 0.077281 |
| ABA 1h | 4500.28 | 1563.34 | 0.464385 | 0.124802 |
| ACC 1h | 20062 | 4235.26 | 1.89411 | 0.309026 |
| BABA 1h | 14441.5 | 917.808 | 1.51933 | 0.090939 |
| Chitin 1h | 14184.3 | 2795.8 | 1.54396 | 0.402695 |
| Epi 1h | 12190 | 876.143 | 1.42662 | 0.0823671 |
| SA 1h | 19664 | 9289.53 | 1.58749 | 0.819841 |
| Me-JA 1h | 34451.9 | 2595.75 | 4.27109 | 0.246592 |
| Control 6h | 13608.8 | 5280.49 | 1.202 | 0.379229 |
| ABA 6h | 1300.89 | 223.432 | 0.120913 | 0.0185251 |
| ACC 6h | 3876.19 | 748.986 | 0.330102 | 0.0538404 |
| BABA 6h | 7644.47 | 525.78 | 0.692157 | 0.0433804 |
| Chitin 6h | 9586.08 | 2283.37 | 0.868767 | 0.219733 |
| Epi 6h | 8062.33 | 2575.64 | 0.62656 | 0.372348 |
| SA 6h | 3673.51 | 598.759 | 0.368289 | 0.0752816 |
| Me-JA 6h | 11910.3 | 1973.1 | 1.17962 | 0.128497 |
Source Transcript PGSC0003DMT400067600 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G43710.1 | +1 | 0.0 | 530 | 263/464 (57%) | Pyridoxal phosphate (PLP)-dependent transferases superfamily protein | chr1:16486534-16488298 REVERSE LENGTH=482 |
