Probe CUST_33229_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33229_PI426222305 | JHI_St_60k_v1 | DMT400067586 | TAAGGTGTTTGTGCATCCAAAGTTCCACTCTCCTAGCAAACCCGATCAATACAGAAAAAA |
All Microarray Probes Designed to Gene DMG400026277
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33229_PI426222305 | JHI_St_60k_v1 | DMT400067586 | TAAGGTGTTTGTGCATCCAAAGTTCCACTCTCCTAGCAAACCCGATCAATACAGAAAAAA |
| CUST_33174_PI426222305 | JHI_St_60k_v1 | DMT400067587 | CCACGTGGGCACCTATATATCTTTAAAATACACTATTAAATAGTGCAGGGGTAATAGATA |
Microarray Signals from CUST_33229_PI426222305
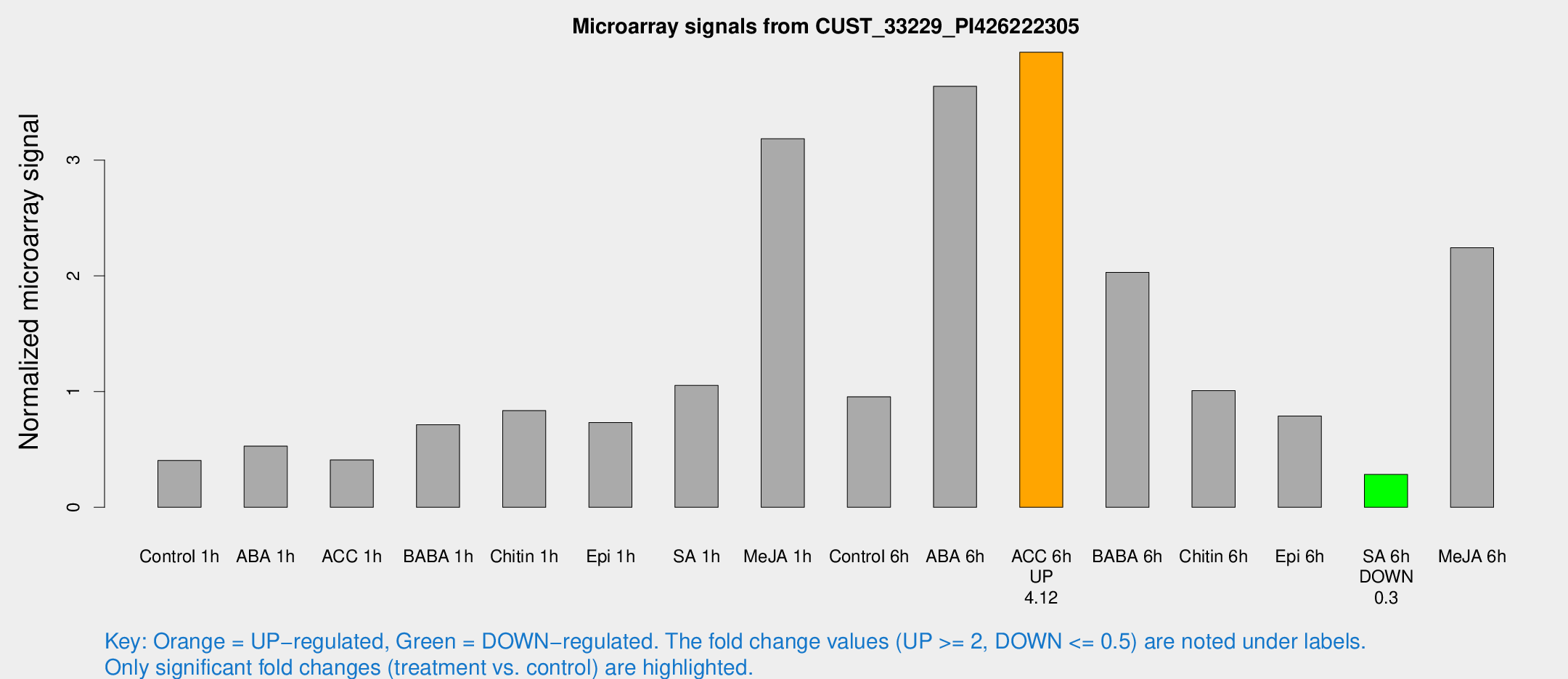
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 50.3951 | 8.07757 | 0.405194 | 0.0801128 |
| ABA 1h | 64.5055 | 20.6323 | 0.528802 | 0.200183 |
| ACC 1h | 62.0157 | 22.4281 | 0.40896 | 0.196817 |
| BABA 1h | 120.798 | 50.963 | 0.713953 | 0.600873 |
| Chitin 1h | 91.6897 | 9.87829 | 0.835294 | 0.0594452 |
| Epi 1h | 76.8367 | 5.74916 | 0.732609 | 0.0549776 |
| SA 1h | 132.007 | 12.886 | 1.05338 | 0.0675003 |
| Me-JA 1h | 356.117 | 106.438 | 3.18475 | 1.0517 |
| Control 6h | 135.25 | 44.5438 | 0.955079 | 0.349147 |
| ABA 6h | 547.555 | 188.145 | 3.638 | 1.52855 |
| ACC 6h | 557.068 | 83.1457 | 3.93183 | 0.886507 |
| BABA 6h | 339.846 | 161.144 | 2.0294 | 1.00806 |
| Chitin 6h | 130.834 | 12.2273 | 1.00842 | 0.129697 |
| Epi 6h | 112.455 | 25.2714 | 0.788915 | 0.173535 |
| SA 6h | 35.1039 | 6.91271 | 0.283453 | 0.0546703 |
| Me-JA 6h | 290.989 | 81.2802 | 2.24225 | 0.745748 |
Source Transcript PGSC0003DMT400067586 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G36690.1 | +1 | 3e-73 | 239 | 120/307 (39%) | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | chr2:15379930-15381987 FORWARD LENGTH=366 |
