Probe CUST_32665_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_32665_PI426222305 | JHI_St_60k_v1 | DMT400047322 | ACCATAATTGAAGTAGATCATTTGCGGATGAAAGACATGAAGATGGTAAATGGAGACGTT |
All Microarray Probes Designed to Gene DMG400018381
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_32735_PI426222305 | JHI_St_60k_v1 | DMT400047321 | ATGATTTTCGTGGATGGCCGGGTTTTTCAGATCATTTGCGGATGAAAGACATGAAGATGG |
| CUST_32665_PI426222305 | JHI_St_60k_v1 | DMT400047322 | ACCATAATTGAAGTAGATCATTTGCGGATGAAAGACATGAAGATGGTAAATGGAGACGTT |
Microarray Signals from CUST_32665_PI426222305
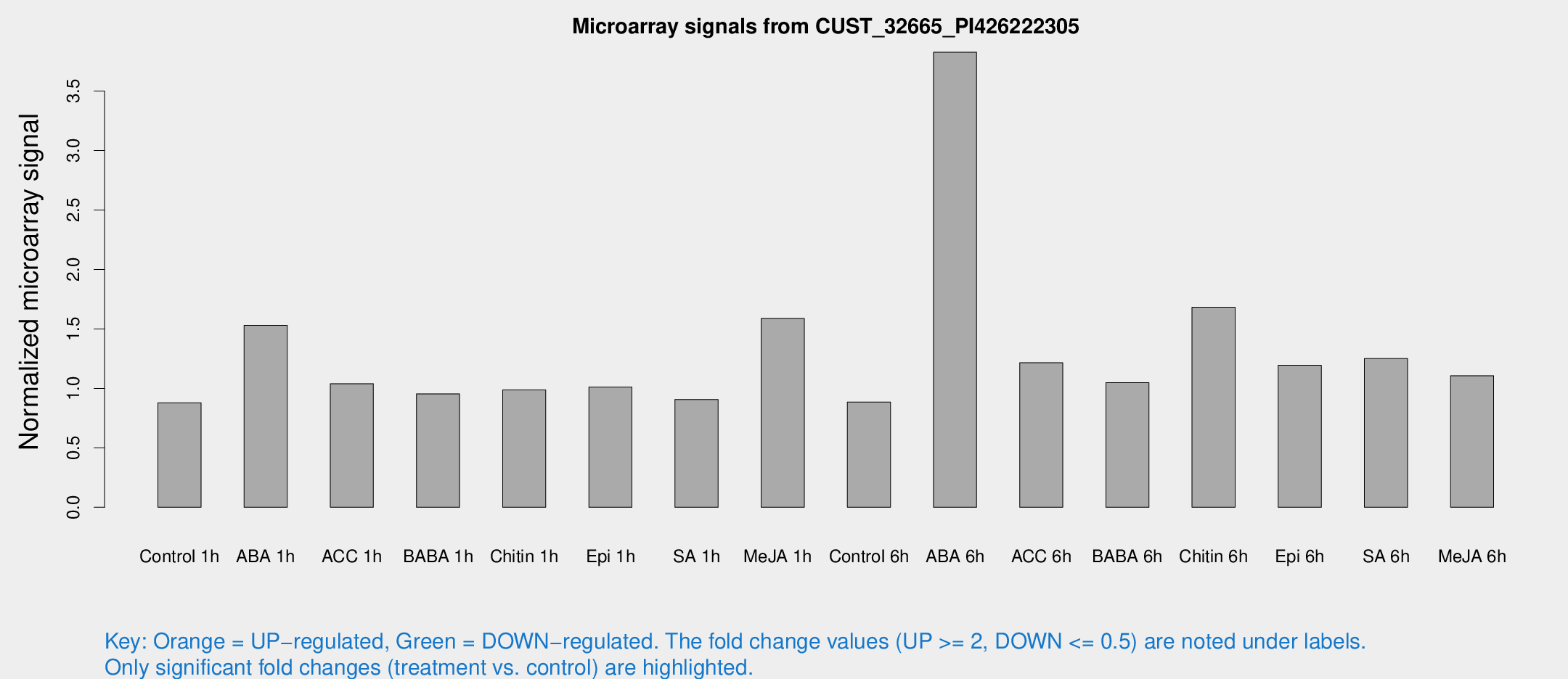
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.09581 | 2.95461 | 0.878393 | 0.501502 |
| ABA 1h | 9.64967 | 4.65047 | 1.52952 | 0.664995 |
| ACC 1h | 6.3389 | 3.50374 | 1.03941 | 0.567592 |
| BABA 1h | 5.41885 | 3.14487 | 0.95444 | 0.553097 |
| Chitin 1h | 5.06125 | 2.94688 | 0.987867 | 0.552875 |
| Epi 1h | 5.02452 | 2.91768 | 1.01185 | 0.571557 |
| SA 1h | 5.50722 | 3.0108 | 0.906668 | 0.499356 |
| Me-JA 1h | 7.86994 | 3.0834 | 1.5876 | 0.66071 |
| Control 6h | 5.22048 | 3.02365 | 0.885461 | 0.512766 |
| ABA 6h | 24.1607 | 3.45588 | 3.82705 | 0.554552 |
| ACC 6h | 8.73057 | 3.72698 | 1.21573 | 0.579656 |
| BABA 6h | 7.05117 | 3.43306 | 1.04861 | 0.533808 |
| Chitin 6h | 12.0457 | 4.38287 | 1.6832 | 0.64336 |
| Epi 6h | 8.66069 | 3.60015 | 1.19401 | 0.559588 |
| SA 6h | 7.71716 | 3.28034 | 1.25047 | 0.605575 |
| Me-JA 6h | 6.86919 | 3.06902 | 1.10674 | 0.548284 |
Source Transcript PGSC0003DMT400047322 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G01180.1 | +1 | 3e-82 | 254 | 127/229 (55%) | S-adenosyl-L-methionine-dependent methyltransferases superfamily protein | chr1:75633-76556 FORWARD LENGTH=307 |
