Probe CUST_32735_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_32735_PI426222305 | JHI_St_60k_v1 | DMT400047321 | ATGATTTTCGTGGATGGCCGGGTTTTTCAGATCATTTGCGGATGAAAGACATGAAGATGG |
All Microarray Probes Designed to Gene DMG400018381
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_32735_PI426222305 | JHI_St_60k_v1 | DMT400047321 | ATGATTTTCGTGGATGGCCGGGTTTTTCAGATCATTTGCGGATGAAAGACATGAAGATGG |
| CUST_32665_PI426222305 | JHI_St_60k_v1 | DMT400047322 | ACCATAATTGAAGTAGATCATTTGCGGATGAAAGACATGAAGATGGTAAATGGAGACGTT |
Microarray Signals from CUST_32735_PI426222305
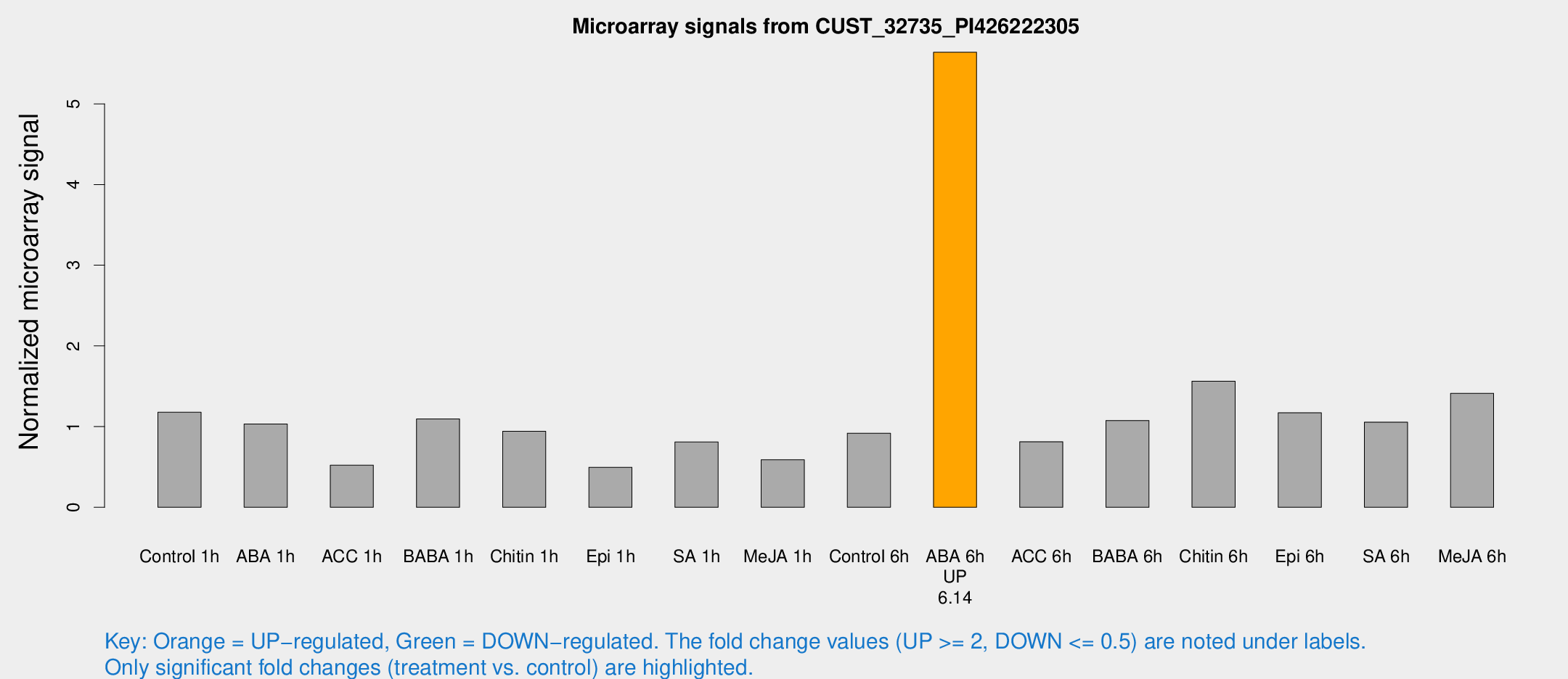
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 20.1008 | 4.07544 | 1.17777 | 0.225631 |
| ABA 1h | 18.5687 | 7.22434 | 1.03307 | 0.526773 |
| ACC 1h | 10.1471 | 4.04038 | 0.521922 | 0.244987 |
| BABA 1h | 17.8086 | 3.72318 | 1.09523 | 0.244721 |
| Chitin 1h | 16.5884 | 6.55695 | 0.941838 | 0.434802 |
| Epi 1h | 7.18669 | 3.37358 | 0.496098 | 0.239323 |
| SA 1h | 16.4042 | 6.07991 | 0.808774 | 0.392237 |
| Me-JA 1h | 8.8173 | 3.51705 | 0.588481 | 0.282905 |
| Control 6h | 15.6562 | 3.63273 | 0.91879 | 0.26671 |
| ABA 6h | 101.57 | 17.2965 | 5.63956 | 0.8773 |
| ACC 6h | 15.2891 | 4.36695 | 0.812637 | 0.230619 |
| BABA 6h | 20.2795 | 4.08139 | 1.07602 | 0.225053 |
| Chitin 6h | 27.3675 | 4.17931 | 1.5618 | 0.240456 |
| Epi 6h | 22.9268 | 5.04008 | 1.17148 | 0.380879 |
| SA 6h | 18.6468 | 5.10578 | 1.05548 | 0.273303 |
| Me-JA 6h | 31.3055 | 15.7034 | 1.41262 | 0.852561 |
Source Transcript PGSC0003DMT400047321 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G01180.1 | +1 | 5e-113 | 335 | 157/231 (68%) | S-adenosyl-L-methionine-dependent methyltransferases superfamily protein | chr1:75633-76556 FORWARD LENGTH=307 |
