Probe CUST_29774_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29774_PI426222305 | JHI_St_60k_v1 | DMT400002644 | ATGCATTAGGGGATTGTTTCTCTGAAATCAACACAAATCCAGGCTACGTCGATGAGGAAG |
All Microarray Probes Designed to Gene DMG400001015
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29769_PI426222305 | JHI_St_60k_v1 | DMT400002645 | CTTCCAGTACCTTACTAAACACAAATGGGCTACCAGATTGTTGTTCTTTGTATTGTATGT |
| CUST_29774_PI426222305 | JHI_St_60k_v1 | DMT400002644 | ATGCATTAGGGGATTGTTTCTCTGAAATCAACACAAATCCAGGCTACGTCGATGAGGAAG |
Microarray Signals from CUST_29774_PI426222305
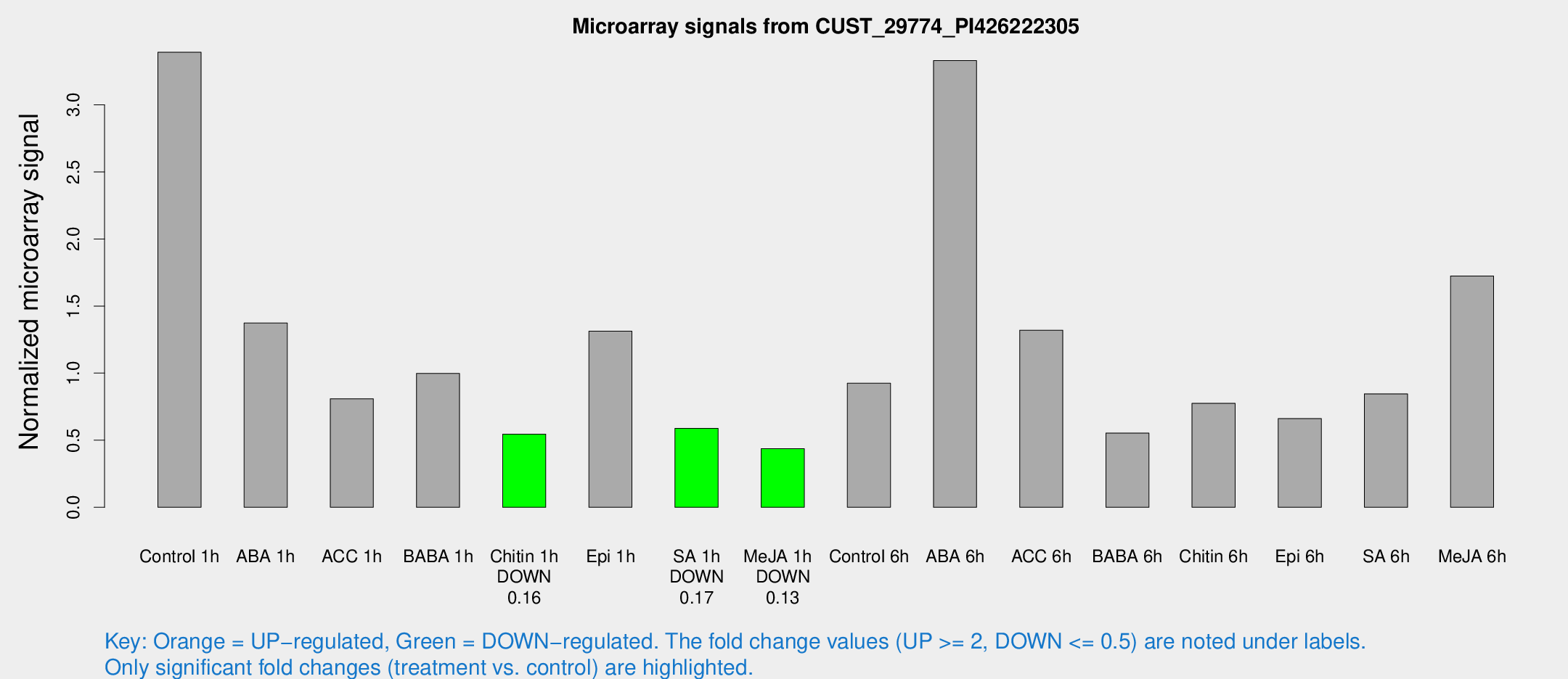
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 114.065 | 16.6848 | 3.39158 | 0.266513 |
| ABA 1h | 43.8534 | 11.4403 | 1.37358 | 0.381179 |
| ACC 1h | 41.0798 | 19.0309 | 0.809351 | 0.780963 |
| BABA 1h | 41.2855 | 16.6161 | 0.997356 | 0.563991 |
| Chitin 1h | 16.2421 | 3.3084 | 0.545634 | 0.112088 |
| Epi 1h | 42.6397 | 13.3329 | 1.31256 | 0.531802 |
| SA 1h | 23.9208 | 8.6494 | 0.58793 | 0.270644 |
| Me-JA 1h | 12.9546 | 4.04092 | 0.436378 | 0.134122 |
| Control 6h | 47.1325 | 29.9624 | 0.924573 | 0.873081 |
| ABA 6h | 123.406 | 26.3695 | 3.32888 | 0.693234 |
| ACC 6h | 54.6819 | 16.8755 | 1.31982 | 0.222993 |
| BABA 6h | 23.947 | 9.75622 | 0.553231 | 0.231157 |
| Chitin 6h | 30.937 | 9.56332 | 0.775228 | 0.341613 |
| Epi 6h | 26.8401 | 6.92877 | 0.66097 | 0.174001 |
| SA 6h | 34.3646 | 16.6534 | 0.845727 | 0.366631 |
| Me-JA 6h | 58.0035 | 9.44234 | 1.72427 | 0.159105 |
Source Transcript PGSC0003DMT400002644 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G73010.1 | +1 | 1e-94 | 281 | 141/220 (64%) | phosphate starvation-induced gene 2 | chr1:27464780-27466180 REVERSE LENGTH=295 |
