Probe CUST_29769_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29769_PI426222305 | JHI_St_60k_v1 | DMT400002645 | CTTCCAGTACCTTACTAAACACAAATGGGCTACCAGATTGTTGTTCTTTGTATTGTATGT |
All Microarray Probes Designed to Gene DMG400001015
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29769_PI426222305 | JHI_St_60k_v1 | DMT400002645 | CTTCCAGTACCTTACTAAACACAAATGGGCTACCAGATTGTTGTTCTTTGTATTGTATGT |
| CUST_29774_PI426222305 | JHI_St_60k_v1 | DMT400002644 | ATGCATTAGGGGATTGTTTCTCTGAAATCAACACAAATCCAGGCTACGTCGATGAGGAAG |
Microarray Signals from CUST_29769_PI426222305
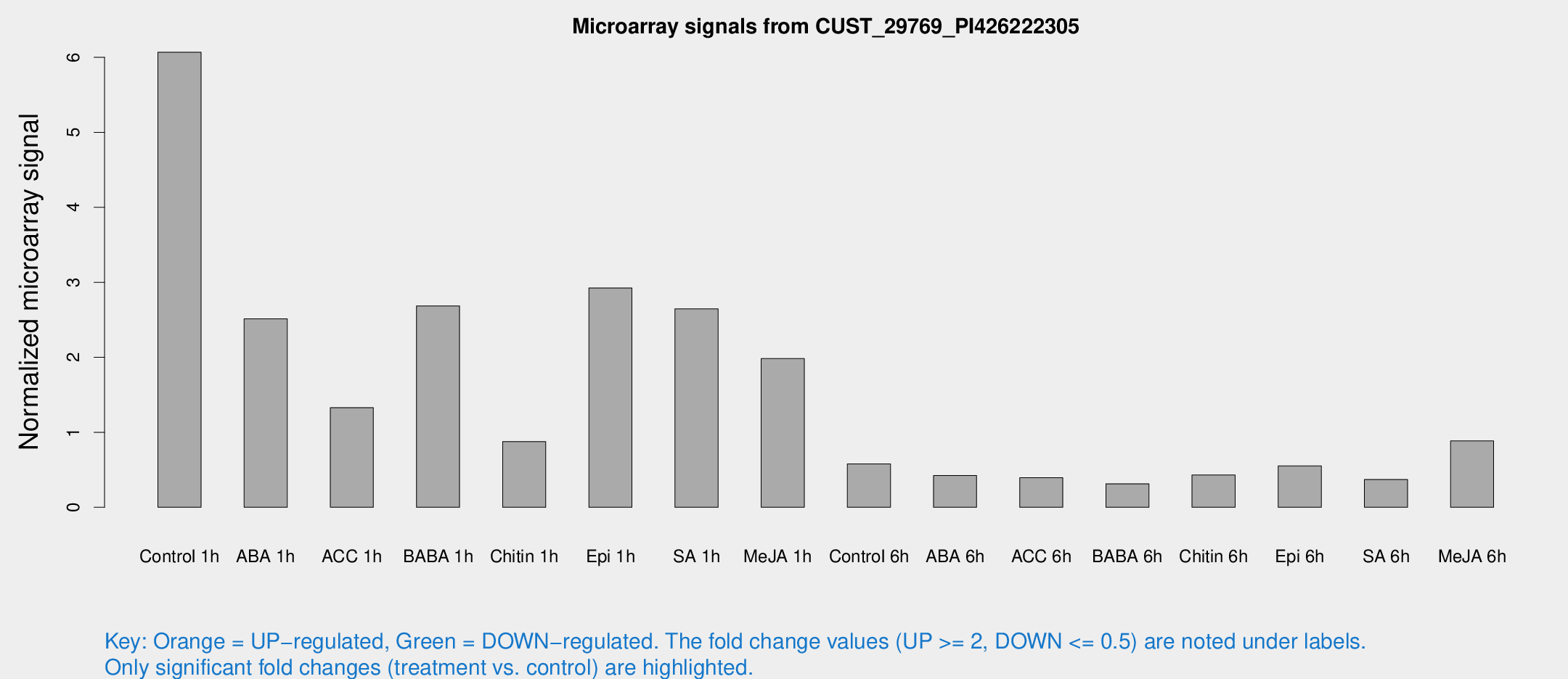
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 329.799 | 166.941 | 6.06801 | 3.27749 |
| ABA 1h | 100.853 | 28.035 | 2.51216 | 0.682381 |
| ACC 1h | 115.177 | 68.1869 | 1.32863 | 2.75006 |
| BABA 1h | 138.997 | 65.3835 | 2.68545 | 1.29339 |
| Chitin 1h | 52.2993 | 23.6206 | 0.876956 | 1.30535 |
| Epi 1h | 136.898 | 52.2914 | 2.92396 | 2.04785 |
| SA 1h | 114.563 | 7.51886 | 2.64509 | 0.173554 |
| Me-JA 1h | 76.7355 | 25.4038 | 1.98265 | 0.69076 |
| Control 6h | 36.3201 | 18.2507 | 0.579462 | 0.700956 |
| ABA 6h | 24.3133 | 9.24816 | 0.423851 | 0.275049 |
| ACC 6h | 23.3419 | 10.7537 | 0.394875 | 0.263976 |
| BABA 6h | 14.8526 | 4.2275 | 0.31315 | 0.0919083 |
| Chitin 6h | 24.3774 | 9.36222 | 0.430689 | 0.297687 |
| Epi 6h | 26.8964 | 4.56668 | 0.552048 | 0.0962346 |
| SA 6h | 16.7579 | 5.01902 | 0.371406 | 0.16598 |
| Me-JA 6h | 45.4502 | 20.7607 | 0.88573 | 0.356382 |
Source Transcript PGSC0003DMT400002645 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G73010.1 | +1 | 3e-49 | 164 | 74/119 (62%) | phosphate starvation-induced gene 2 | chr1:27464780-27466180 REVERSE LENGTH=295 |
