Probe CUST_2927_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2927_PI426222305 | JHI_St_60k_v1 | DMT400000352 | CTGTTGTTTTTTCCCTGATCCTTTCTAGGAGGGGGTTTCATGTATTATAAATTCTAGTTT |
All Microarray Probes Designed to Gene DMG400000123
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3165_PI426222305 | JHI_St_60k_v1 | DMT400000353 | CTGTTGTTTTTTCCCTGATCCTTTCTAGGAGGGGGTTTCATGTATTATAAATTCTAGTTT |
| CUST_2927_PI426222305 | JHI_St_60k_v1 | DMT400000352 | CTGTTGTTTTTTCCCTGATCCTTTCTAGGAGGGGGTTTCATGTATTATAAATTCTAGTTT |
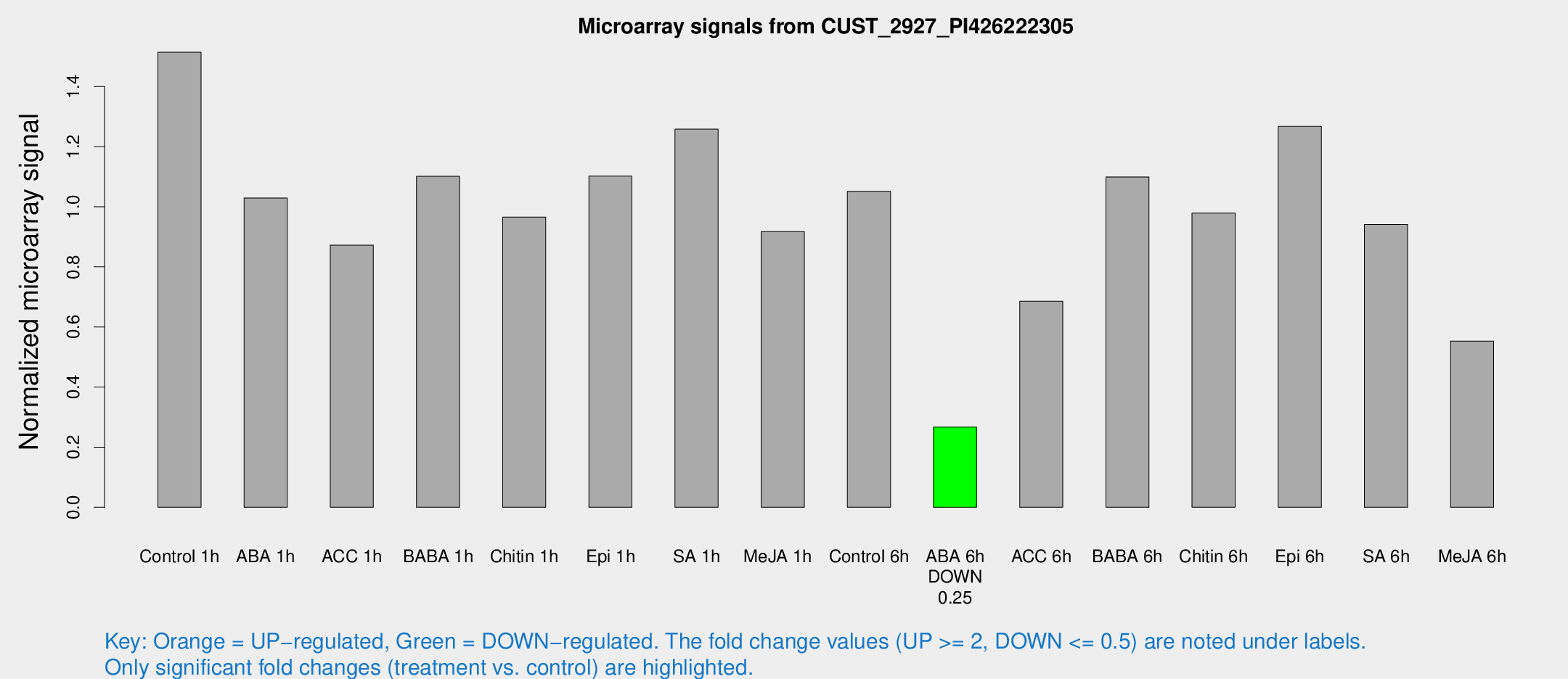
Microarray Signals from CUST_2927_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1501.82 | 374.017 | 1.51412 | 0.273966 |
| ABA 1h | 856.522 | 72.4913 | 1.02921 | 0.0752171 |
| ACC 1h | 859.88 | 132.363 | 0.87235 | 0.0806321 |
| BABA 1h | 1037.46 | 202.199 | 1.10114 | 0.129228 |
| Chitin 1h | 819.828 | 84.5423 | 0.965663 | 0.0908506 |
| Epi 1h | 895.943 | 68.942 | 1.10193 | 0.0637388 |
| SA 1h | 1213.05 | 86.3757 | 1.25805 | 0.0727122 |
| Me-JA 1h | 707.402 | 76.5836 | 0.917378 | 0.059934 |
| Control 6h | 1046.6 | 232.379 | 1.05158 | 0.175359 |
| ABA 6h | 266.875 | 21.8601 | 0.267019 | 0.015795 |
| ACC 6h | 742.087 | 72.2083 | 0.685518 | 0.0655132 |
| BABA 6h | 1156.82 | 90.8298 | 1.09918 | 0.0635584 |
| Chitin 6h | 983.126 | 96.9291 | 0.979212 | 0.0652182 |
| Epi 6h | 1346.6 | 125.198 | 1.26704 | 0.0794157 |
| SA 6h | 884.021 | 111.96 | 0.940826 | 0.0544639 |
| Me-JA 6h | 594.167 | 206.08 | 0.553194 | 0.185456 |
Source Transcript PGSC0003DMT400000352 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G00900.1 | +3 | 0.0 | 1607 | 814/1051 (77%) | ER-type Ca2+-ATPase 2 | chr4:382690-386226 REVERSE LENGTH=1054 |
