Probe CUST_3165_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3165_PI426222305 | JHI_St_60k_v1 | DMT400000353 | CTGTTGTTTTTTCCCTGATCCTTTCTAGGAGGGGGTTTCATGTATTATAAATTCTAGTTT |
All Microarray Probes Designed to Gene DMG400000123
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3165_PI426222305 | JHI_St_60k_v1 | DMT400000353 | CTGTTGTTTTTTCCCTGATCCTTTCTAGGAGGGGGTTTCATGTATTATAAATTCTAGTTT |
| CUST_2927_PI426222305 | JHI_St_60k_v1 | DMT400000352 | CTGTTGTTTTTTCCCTGATCCTTTCTAGGAGGGGGTTTCATGTATTATAAATTCTAGTTT |
Microarray Signals from CUST_3165_PI426222305
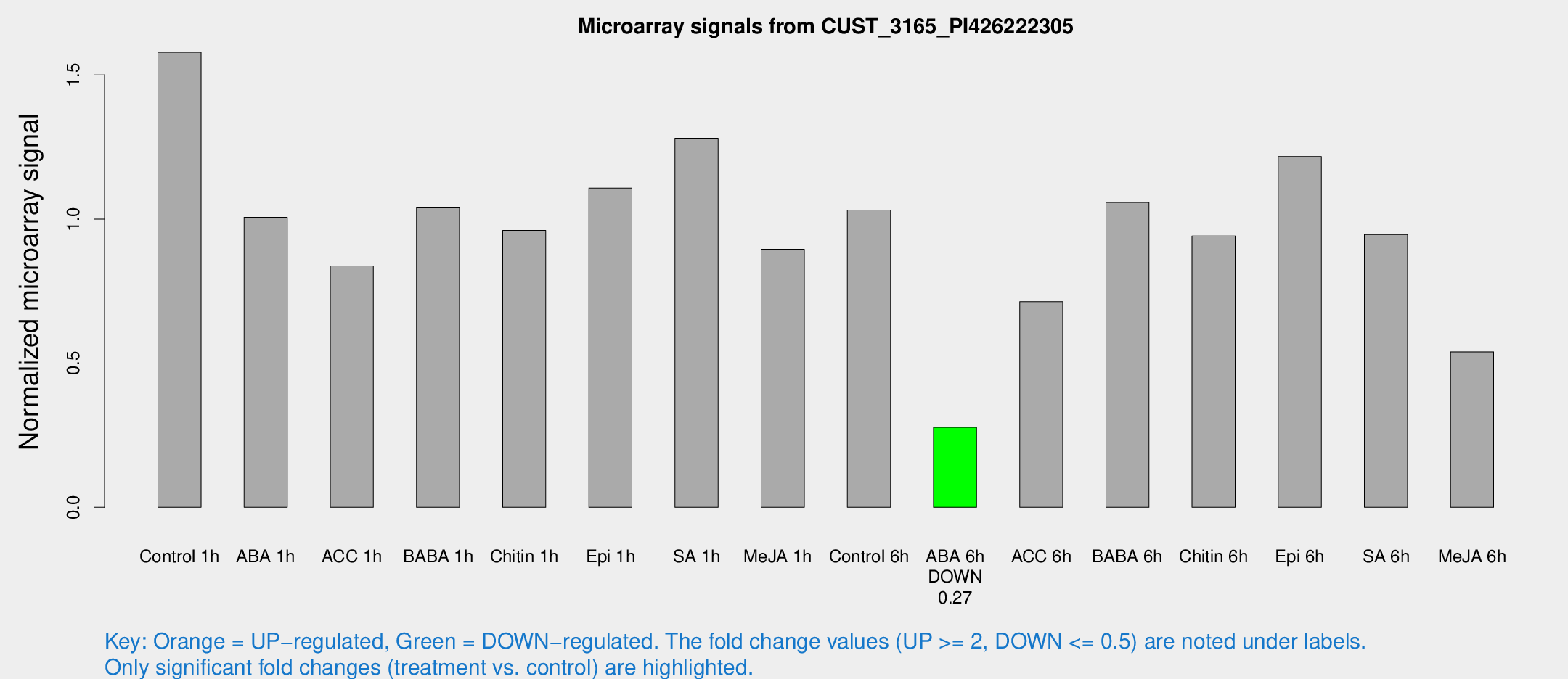
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1536.78 | 321.806 | 1.57851 | 0.219616 |
| ABA 1h | 843.649 | 100.243 | 1.00636 | 0.124544 |
| ACC 1h | 844.718 | 172.302 | 0.837629 | 0.1327 |
| BABA 1h | 991.05 | 226.307 | 1.03909 | 0.160254 |
| Chitin 1h | 826.295 | 133.409 | 0.960585 | 0.154753 |
| Epi 1h | 903.411 | 91.7167 | 1.10725 | 0.0796437 |
| SA 1h | 1232.07 | 71.4598 | 1.28042 | 0.0740183 |
| Me-JA 1h | 685.77 | 49.2748 | 0.895292 | 0.0632676 |
| Control 6h | 1016.24 | 210.128 | 1.03115 | 0.153866 |
| ABA 6h | 277.228 | 22.0693 | 0.277633 | 0.0165211 |
| ACC 6h | 777.742 | 103.823 | 0.713425 | 0.0414003 |
| BABA 6h | 1121.44 | 134.875 | 1.05745 | 0.104039 |
| Chitin 6h | 942.527 | 78.8988 | 0.94157 | 0.0545246 |
| Epi 6h | 1287.81 | 93.9802 | 1.21708 | 0.088813 |
| SA 6h | 884.779 | 97.7755 | 0.946537 | 0.0557209 |
| Me-JA 6h | 574.85 | 205.462 | 0.539617 | 0.167078 |
Source Transcript PGSC0003DMT400000353 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G00900.1 | +3 | 0.0 | 1337 | 677/865 (78%) | ER-type Ca2+-ATPase 2 | chr4:382690-386226 REVERSE LENGTH=1054 |
