Probe CUST_2887_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2887_PI426222305 | JHI_St_60k_v1 | DMT400000220 | AGATCTCTTACACTCTCAAATGCTCCGGCAGGGTTATAGTATCAGGAACAATTCCAGATC |
All Microarray Probes Designed to Gene DMG400000066
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2887_PI426222305 | JHI_St_60k_v1 | DMT400000220 | AGATCTCTTACACTCTCAAATGCTCCGGCAGGGTTATAGTATCAGGAACAATTCCAGATC |
| CUST_3229_PI426222305 | JHI_St_60k_v1 | DMT400000221 | CCAATAAATCTCCTTCCTATATATATAGTCATGTGTGTTAGCATGTATTCCAGATTTGGG |
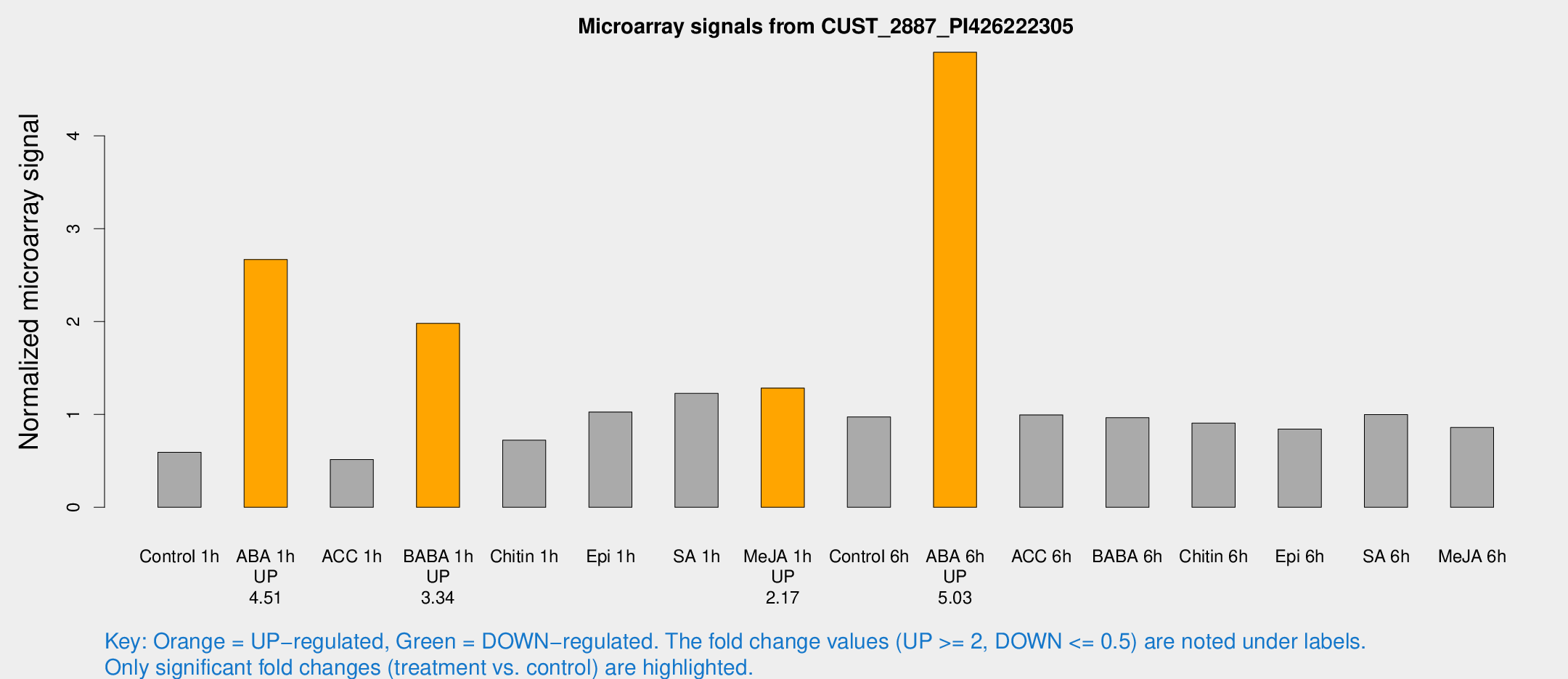
Microarray Signals from CUST_2887_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1720.57 | 128.818 | 0.592041 | 0.0622864 |
| ABA 1h | 7184.66 | 1486.87 | 2.66936 | 0.486051 |
| ACC 1h | 1605.66 | 327.509 | 0.515157 | 0.0819818 |
| BABA 1h | 5545.92 | 423.363 | 1.98019 | 0.362579 |
| Chitin 1h | 1919.83 | 288.76 | 0.722663 | 0.0507977 |
| Epi 1h | 2670.58 | 528.224 | 1.02653 | 0.239028 |
| SA 1h | 4016.71 | 1168.13 | 1.2263 | 0.342218 |
| Me-JA 1h | 3139 | 558.824 | 1.28394 | 0.148919 |
| Control 6h | 3040.02 | 764.634 | 0.973885 | 0.202747 |
| ABA 6h | 15435.2 | 2297.44 | 4.90088 | 0.574017 |
| ACC 6h | 3431.5 | 709.246 | 0.995579 | 0.0692816 |
| BABA 6h | 3156.22 | 339.161 | 0.965106 | 0.0887989 |
| Chitin 6h | 2804.79 | 204.88 | 0.907008 | 0.0911648 |
| Epi 6h | 2785.38 | 331.023 | 0.843351 | 0.0888161 |
| SA 6h | 2986.66 | 585.927 | 0.999324 | 0.103823 |
| Me-JA 6h | 2518.65 | 319.325 | 0.860416 | 0.0496901 |
Source Transcript PGSC0003DMT400000220 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G01470.1 | +1 | 8e-56 | 180 | 83/151 (55%) | Late embryogenesis abundant protein | chr1:172295-172826 REVERSE LENGTH=151 |
