Probe CUST_3229_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3229_PI426222305 | JHI_St_60k_v1 | DMT400000221 | CCAATAAATCTCCTTCCTATATATATAGTCATGTGTGTTAGCATGTATTCCAGATTTGGG |
All Microarray Probes Designed to Gene DMG400000066
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2887_PI426222305 | JHI_St_60k_v1 | DMT400000220 | AGATCTCTTACACTCTCAAATGCTCCGGCAGGGTTATAGTATCAGGAACAATTCCAGATC |
| CUST_3229_PI426222305 | JHI_St_60k_v1 | DMT400000221 | CCAATAAATCTCCTTCCTATATATATAGTCATGTGTGTTAGCATGTATTCCAGATTTGGG |
Microarray Signals from CUST_3229_PI426222305
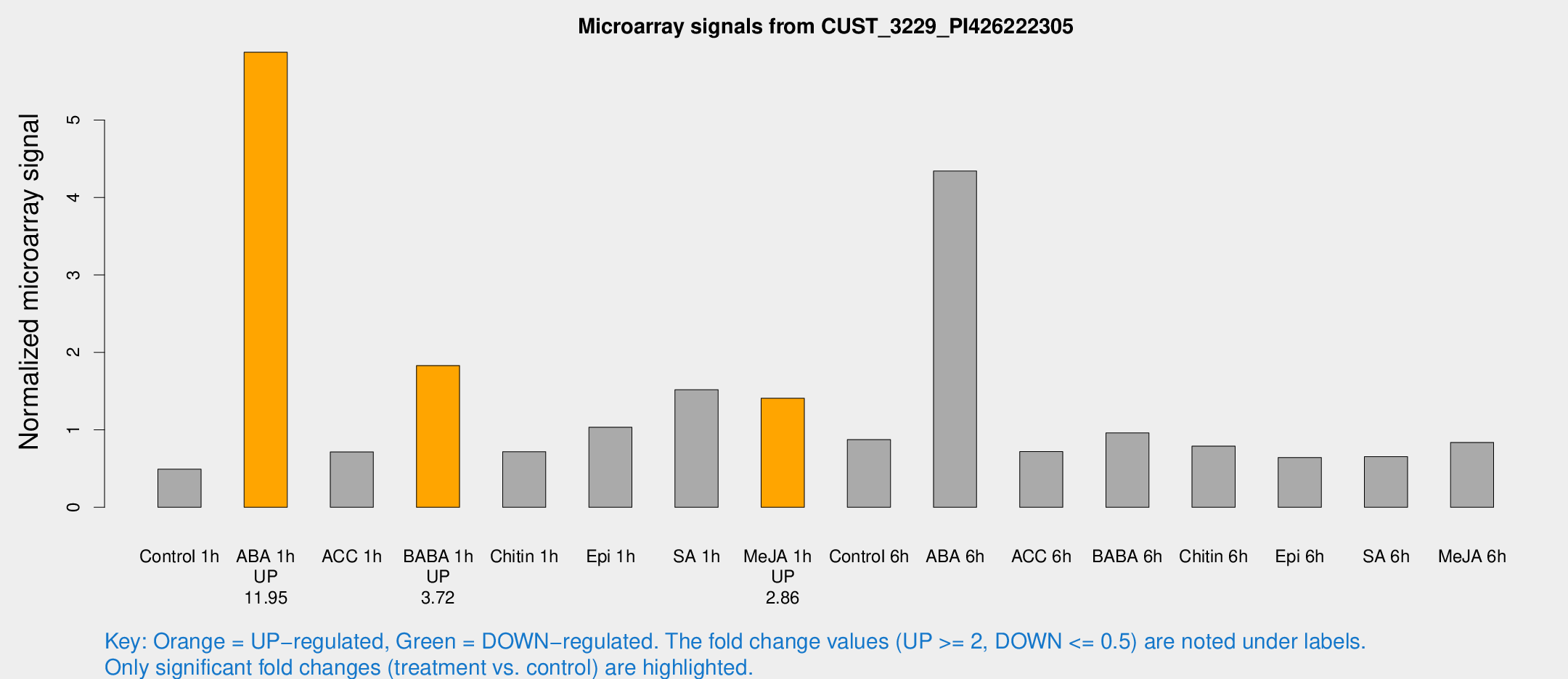
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11.6876 | 3.61778 | 0.491709 | 0.15411 |
| ABA 1h | 130.639 | 29.2816 | 5.87441 | 1.28473 |
| ACC 1h | 19.5448 | 5.71758 | 0.713817 | 0.230998 |
| BABA 1h | 41.7154 | 4.59588 | 1.82938 | 0.219044 |
| Chitin 1h | 15.5192 | 3.80019 | 0.718088 | 0.18914 |
| Epi 1h | 21.9183 | 4.37057 | 1.03383 | 0.215878 |
| SA 1h | 42.9943 | 14.2467 | 1.51869 | 0.621224 |
| Me-JA 1h | 27.9073 | 4.91821 | 1.40754 | 0.205153 |
| Control 6h | 26.4667 | 10.0106 | 0.873752 | 0.48839 |
| ABA 6h | 121.26 | 33.7999 | 4.33992 | 1.4433 |
| ACC 6h | 22.6273 | 7.48081 | 0.719544 | 0.251243 |
| BABA 6h | 26.7036 | 5.52559 | 0.960105 | 0.199592 |
| Chitin 6h | 20.4473 | 4.50648 | 0.789977 | 0.189247 |
| Epi 6h | 17.4133 | 4.57683 | 0.643182 | 0.179784 |
| SA 6h | 17.8656 | 6.69161 | 0.653211 | 0.228942 |
| Me-JA 6h | 25.5865 | 9.90053 | 0.837573 | 0.512627 |
Source Transcript PGSC0003DMT400000221 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G01470.1 | +3 | 3e-41 | 113 | 50/83 (60%) | Late embryogenesis abundant protein | chr1:172295-172826 REVERSE LENGTH=151 |
