Probe CUST_28587_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28587_PI426222305 | JHI_St_60k_v1 | DMT400009899 | CGAGGTAAGAGATAATTTTCGAGTTTAAATTTTATATACTGACCTGACCTGACGTACACT |
All Microarray Probes Designed to Gene DMG400003880
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28587_PI426222305 | JHI_St_60k_v1 | DMT400009899 | CGAGGTAAGAGATAATTTTCGAGTTTAAATTTTATATACTGACCTGACCTGACGTACACT |
| CUST_28527_PI426222305 | JHI_St_60k_v1 | DMT400009900 | CGACAAGACAAACTATTTATCATAAGGATAGGTGTCAAATGAAAGCGCAATGAAAGAACT |
Microarray Signals from CUST_28587_PI426222305
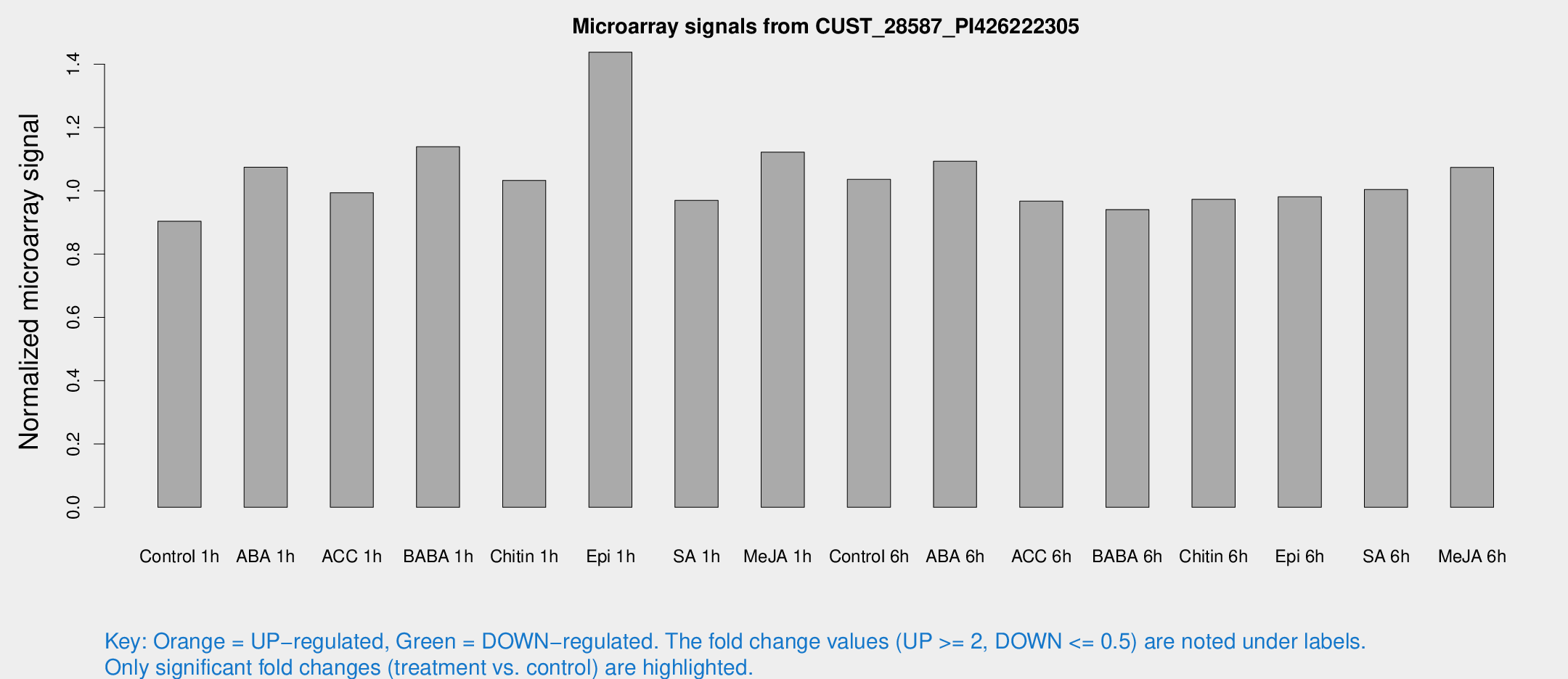
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.35739 | 3.68153 | 0.903455 | 0.523148 |
| ABA 1h | 6.69318 | 3.64742 | 1.07459 | 0.586596 |
| ACC 1h | 7.2196 | 4.20544 | 0.993937 | 0.576119 |
| BABA 1h | 7.82252 | 3.98677 | 1.13924 | 0.596395 |
| Chitin 1h | 6.55559 | 3.80949 | 1.03283 | 0.598344 |
| Epi 1h | 9.07695 | 3.71883 | 1.43769 | 0.637046 |
| SA 1h | 7.08632 | 3.70174 | 0.969764 | 0.516021 |
| Me-JA 1h | 6.465 | 3.75465 | 1.12215 | 0.649742 |
| Control 6h | 7.3631 | 3.92566 | 1.03602 | 0.55989 |
| ABA 6h | 8.40123 | 4.07058 | 1.09333 | 0.556763 |
| ACC 6h | 8.01736 | 4.77647 | 0.967051 | 0.560366 |
| BABA 6h | 7.43946 | 4.32655 | 0.940479 | 0.544613 |
| Chitin 6h | 7.28993 | 4.22336 | 0.972989 | 0.563531 |
| Epi 6h | 7.83935 | 4.57739 | 0.980916 | 0.568675 |
| SA 6h | 6.99107 | 4.05008 | 1.00417 | 0.581575 |
| Me-JA 6h | 7.72669 | 3.81485 | 1.07386 | 0.554312 |
Source Transcript PGSC0003DMT400009899 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G36690.1 | +2 | 9e-121 | 356 | 178/307 (58%) | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | chr2:15379930-15381987 FORWARD LENGTH=366 |
