Probe CUST_28527_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28527_PI426222305 | JHI_St_60k_v1 | DMT400009900 | CGACAAGACAAACTATTTATCATAAGGATAGGTGTCAAATGAAAGCGCAATGAAAGAACT |
All Microarray Probes Designed to Gene DMG400003880
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28587_PI426222305 | JHI_St_60k_v1 | DMT400009899 | CGAGGTAAGAGATAATTTTCGAGTTTAAATTTTATATACTGACCTGACCTGACGTACACT |
| CUST_28527_PI426222305 | JHI_St_60k_v1 | DMT400009900 | CGACAAGACAAACTATTTATCATAAGGATAGGTGTCAAATGAAAGCGCAATGAAAGAACT |
Microarray Signals from CUST_28527_PI426222305
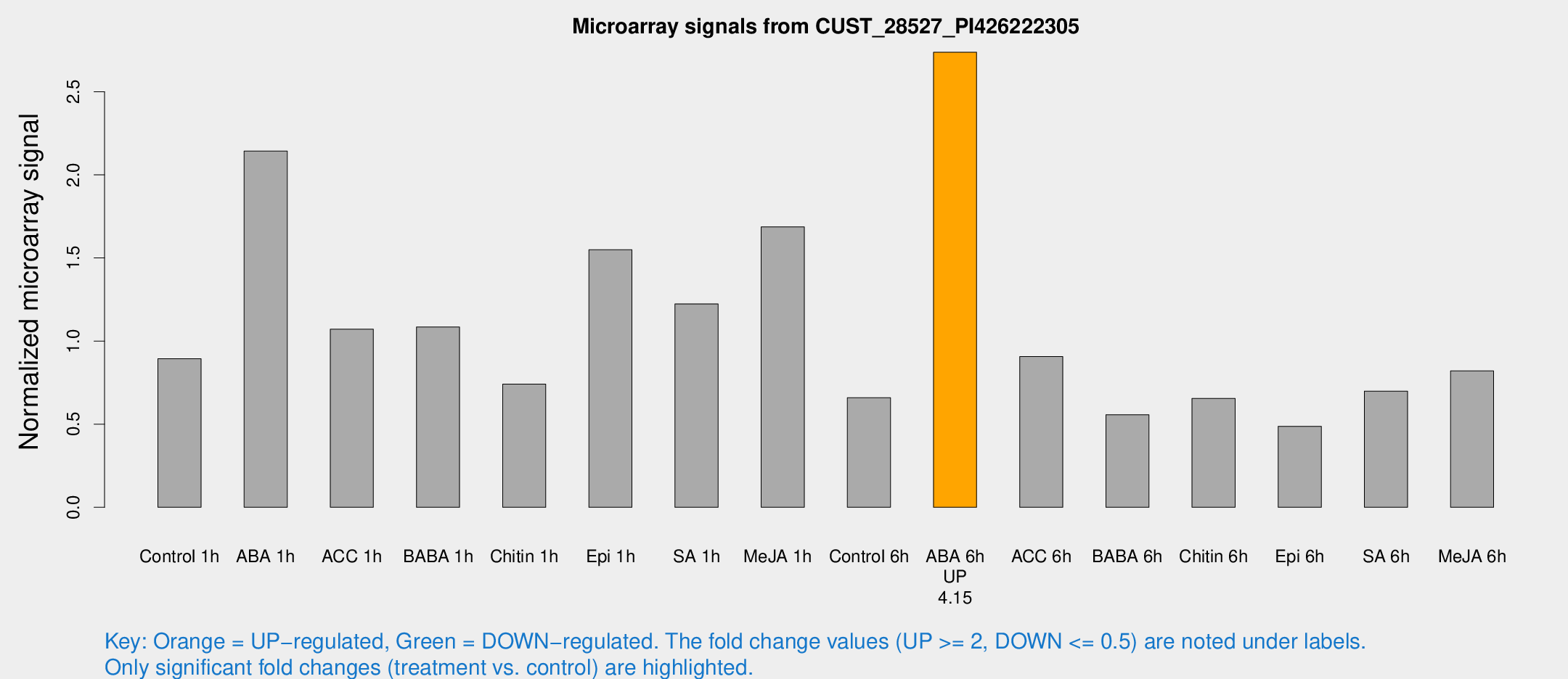
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11.3193 | 3.19325 | 0.89425 | 0.307507 |
| ABA 1h | 22.4534 | 3.33576 | 2.14273 | 0.328228 |
| ACC 1h | 14.5887 | 4.24941 | 1.07165 | 0.366125 |
| BABA 1h | 14.1617 | 4.36802 | 1.0847 | 0.37302 |
| Chitin 1h | 9.286 | 4.13467 | 0.741039 | 0.343111 |
| Epi 1h | 15.9177 | 3.26298 | 1.5491 | 0.327915 |
| SA 1h | 16.8753 | 6.1479 | 1.22366 | 0.39234 |
| Me-JA 1h | 18.785 | 6.36473 | 1.68735 | 0.866767 |
| Control 6h | 8.51389 | 3.4241 | 0.658884 | 0.315644 |
| ABA 6h | 34.9404 | 4.8095 | 2.73761 | 0.324789 |
| ACC 6h | 12.7714 | 4.08641 | 0.906978 | 0.337277 |
| BABA 6h | 7.5816 | 3.81091 | 0.556884 | 0.291577 |
| Chitin 6h | 8.76303 | 3.71188 | 0.655105 | 0.317531 |
| Epi 6h | 6.53508 | 3.8293 | 0.487471 | 0.282652 |
| SA 6h | 8.47973 | 3.56028 | 0.698998 | 0.318975 |
| Me-JA 6h | 10.8323 | 3.7645 | 0.820368 | 0.330606 |
Source Transcript PGSC0003DMT400009900 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G36690.1 | +2 | 1e-157 | 458 | 227/381 (60%) | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | chr2:15379930-15381987 FORWARD LENGTH=366 |
