Probe CUST_28370_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28370_PI426222305 | JHI_St_60k_v1 | DMT400044452 | ATGGATTCATCTAGGGGAATCATGTCAGGAACTATGGATCGGTTCAAAATGATGCTAGTC |
All Microarray Probes Designed to Gene DMG400017253
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28335_PI426222305 | JHI_St_60k_v1 | DMT400044453 | CAGTTAGTATTACCAATGTCTTTGTGCCGGCTTCATTGAACAATAATTGTAAGGGAAATA |
| CUST_28370_PI426222305 | JHI_St_60k_v1 | DMT400044452 | ATGGATTCATCTAGGGGAATCATGTCAGGAACTATGGATCGGTTCAAAATGATGCTAGTC |
Microarray Signals from CUST_28370_PI426222305
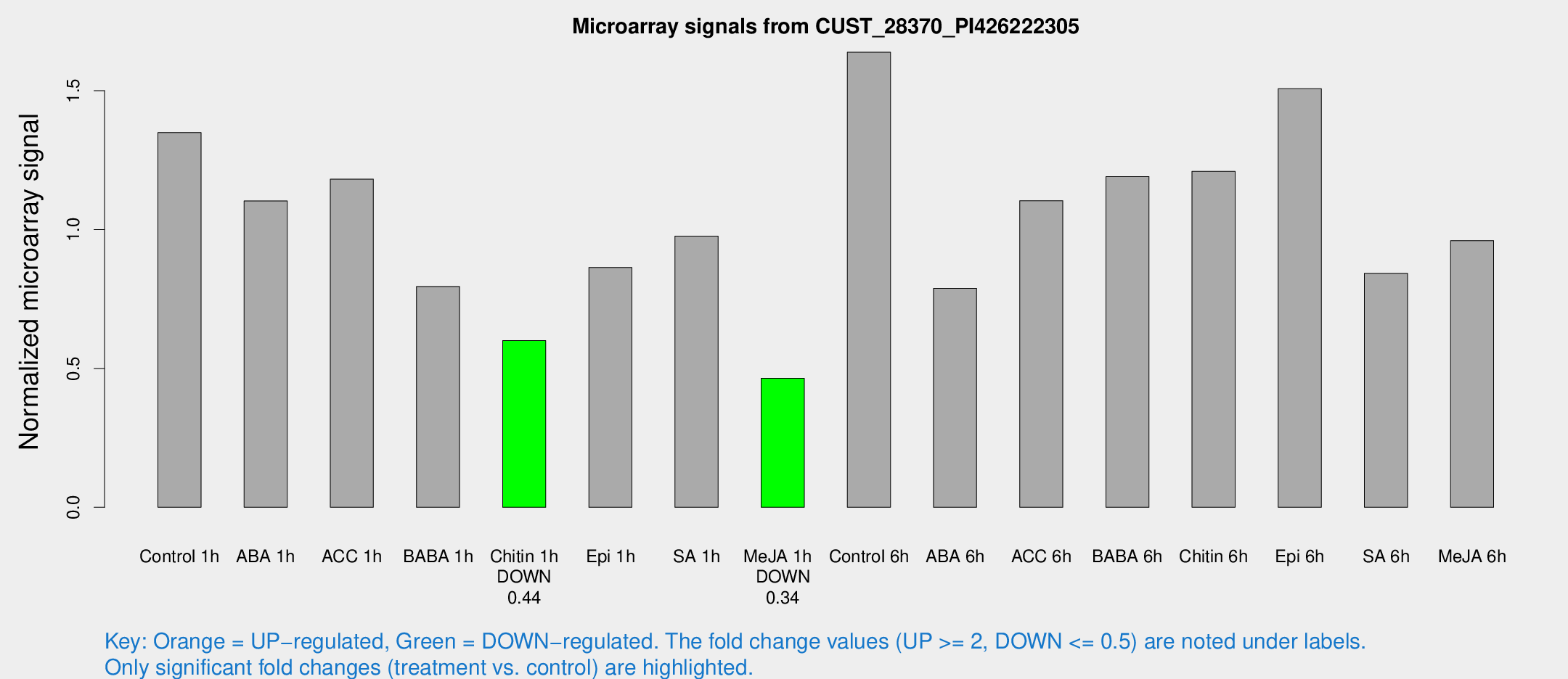
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 114.427 | 12.908 | 1.34934 | 0.0887262 |
| ABA 1h | 81.9677 | 5.83419 | 1.1032 | 0.0784752 |
| ACC 1h | 112.026 | 30.5018 | 1.18153 | 0.313834 |
| BABA 1h | 66.2629 | 11.891 | 0.795061 | 0.0847675 |
| Chitin 1h | 45.9366 | 5.6835 | 0.600084 | 0.0581036 |
| Epi 1h | 64.4037 | 11.3797 | 0.863397 | 0.122726 |
| SA 1h | 85.8517 | 12.8853 | 0.976487 | 0.177181 |
| Me-JA 1h | 31.8181 | 4.09815 | 0.46429 | 0.0595109 |
| Control 6h | 159.811 | 57.1014 | 1.63836 | 0.579773 |
| ABA 6h | 72.3261 | 13.0825 | 0.788333 | 0.179082 |
| ACC 6h | 108.19 | 14.5475 | 1.10374 | 0.123439 |
| BABA 6h | 113.891 | 15.3671 | 1.19066 | 0.133115 |
| Chitin 6h | 111.575 | 21.3811 | 1.20962 | 0.221905 |
| Epi 6h | 148.051 | 30.4782 | 1.50699 | 0.343268 |
| SA 6h | 71.129 | 9.43681 | 0.8423 | 0.0695203 |
| Me-JA 6h | 80.1651 | 5.91263 | 0.960019 | 0.0708416 |
Source Transcript PGSC0003DMT400044452 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G14455.1 | +1 | 7e-39 | 129 | 60/93 (65%) | Target SNARE coiled-coil domain protein | chr4:8310482-8311938 FORWARD LENGTH=130 |
