Probe CUST_28335_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28335_PI426222305 | JHI_St_60k_v1 | DMT400044453 | CAGTTAGTATTACCAATGTCTTTGTGCCGGCTTCATTGAACAATAATTGTAAGGGAAATA |
All Microarray Probes Designed to Gene DMG400017253
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28335_PI426222305 | JHI_St_60k_v1 | DMT400044453 | CAGTTAGTATTACCAATGTCTTTGTGCCGGCTTCATTGAACAATAATTGTAAGGGAAATA |
| CUST_28370_PI426222305 | JHI_St_60k_v1 | DMT400044452 | ATGGATTCATCTAGGGGAATCATGTCAGGAACTATGGATCGGTTCAAAATGATGCTAGTC |
Microarray Signals from CUST_28335_PI426222305
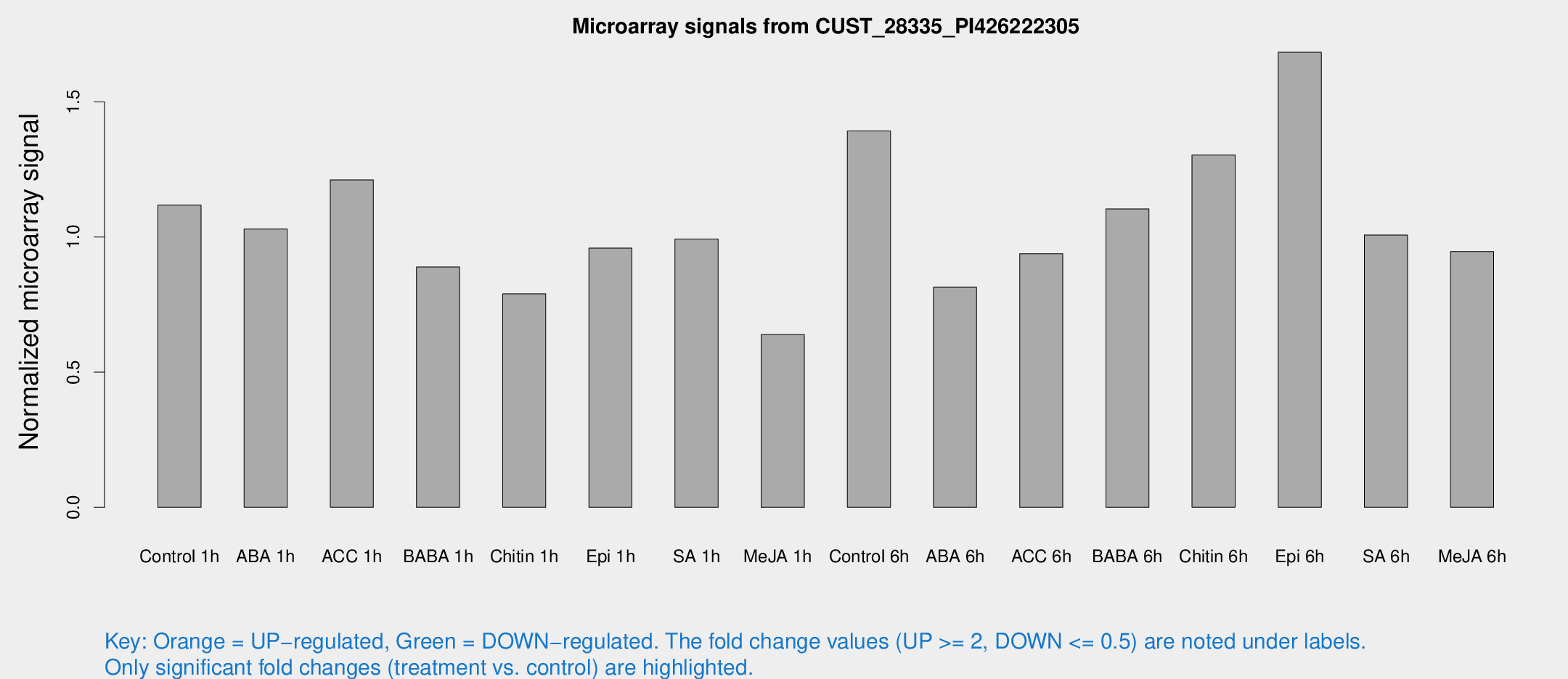
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 148.966 | 41.8505 | 1.11799 | 0.244188 |
| ABA 1h | 112.963 | 7.13898 | 1.02975 | 0.0650715 |
| ACC 1h | 171.661 | 47.7424 | 1.21102 | 0.350166 |
| BABA 1h | 107.099 | 9.54673 | 0.889024 | 0.12194 |
| Chitin 1h | 89.1979 | 9.91988 | 0.790229 | 0.0822306 |
| Epi 1h | 105.406 | 15.5151 | 0.958775 | 0.136291 |
| SA 1h | 132.652 | 26.4284 | 0.992516 | 0.298283 |
| Me-JA 1h | 65.3114 | 6.50795 | 0.638964 | 0.0995759 |
| Control 6h | 190.691 | 51.1295 | 1.39244 | 0.34899 |
| ABA 6h | 108.58 | 11.4755 | 0.814264 | 0.113997 |
| ACC 6h | 134.997 | 15.0227 | 0.938031 | 0.0596532 |
| BABA 6h | 157.4 | 25.7315 | 1.10372 | 0.140304 |
| Chitin 6h | 176.324 | 28.2584 | 1.30301 | 0.197821 |
| Epi 6h | 265.205 | 95.6186 | 1.68361 | 0.644434 |
| SA 6h | 124.426 | 10.3686 | 1.00699 | 0.0643362 |
| Me-JA 6h | 124.478 | 32.7881 | 0.946581 | 0.171188 |
Source Transcript PGSC0003DMT400044453 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G14455.1 | +1 | 1e-55 | 181 | 84/129 (65%) | Target SNARE coiled-coil domain protein | chr4:8310482-8311938 FORWARD LENGTH=130 |
